
MiR-95-3p acts as a prognostic marker and promotes cervical cancer progression by targeting VCAM1
Introduction
Cervical cancer is a relatively common gynecological malignancy (1). This malignancy is caused by a variety of factors, including genetic effects and the human papillomavirus (HPV) viral infection (2). With the development of the HPV vaccination, early diagnostic procedures of cervical dysplasia and cancer can reduce the incidence, morbidity, and mortality of cervical cancer (3). However, advanced cervical cancer patients usually have a high risk of metastasis and a poor prognosis with shorter disease-free survival and overall survival (1). A previous study showed that some factors, such as high parity and poor hygiene, are related to cervical carcinogenesis (4); however, the exact developmental process of cervical cancer remains unknown. Thus, novel prognostic markers and therapeutic targets urgently need to be identified for cervical cancer diagnosis.
Micro ribonucleic acids (miRNAs) are a class of small non-coding RNA molecules that exhibit tissue-specific and developmental-stage-specific expression patterns in a variety of eukaryotes. MiRNAs can induce target messenger RNA degradation or translational repression by binding the complementary sequences in 3'-untranslated regions (3'-UTRs) (5,6). There is emerging evidence that miRNAs are involved in various cellular processes, including cell development, proliferation, and differentiation (6-8). MiRNAs also play important roles in numerous key pathways related to tumor pathogenesis (9). They can function as either carcinogenic factors or tumor suppressors in different tumors (10,11).
There is accumulating evidence that miRNAs are abnormally expressed in cervical cancer tissues and involved in cervical cancer development, progression, metastasis, clinical outcomes, and treatment responses (12-15). MiRNAs can regulate gene expression by targeting multiple molecules to promote or inhibit tumorigenesis (16,17). Wang et al. reported that miR-29a modulates cyclin dependent kinase inhibitor 2A (p16) methylation, which in turn inhibits cervical cancer cell proliferation (18). MiR-155 was shown to promote cervical cancer cell proliferation by suppressing serine/threonine kinase 11 (LKB1) (19). MiR-874 functions as a well-known tumor suppressor in cervical cancer by directly targeting ETS proto-oncogene 1 (ETS1) (20). Thus, miRNAs might be used clinically as diagnostic or prognostic biomarkers and provide treatment strategies for cervical cancer.
However, due to the limited number of patients and different sequencing platforms, most studies have been unable to conduct integrated analyses and there is no recognized standard for the analysis of cervical cancer sequencing data. The Cancer Genome Atlas (TCGA), which is a landmark cancer genomics database, comprises multiple levels of tumor data, including genomic, epigenetic, transcriptomic, proteomic, and clinical data (21). It is a relatively comprehensive public database and has been widely used in various cancer research fields. TCGA contains more than 20,000 tumor and matched normal samples, covers 33 cancer types, and provides massive data that can be used to detect significant genomic changes and discover the biological mechanisms that affect tumor initiation, development, differentiation, invasion, metastasis, and chemosensitivity (22-25).
In this study, we identified miR-95-3p as a poor prognosis-related marker of cervical cancer. We also showed that the upregulation of miR-95-3p induced cervical cancer cell proliferation and migration. Further, we discovered that vascular cell adhesion molecule 1 (VCAM1) is a target gene of miR-95-3p in cervical cancer, and miR-95-3p promotes the malignant behavior of cervical cancer cells by targeting VCAM1. Our findings provide novel insights into specific prognostic biomarkers and potential therapeutic targets for cervical cancer. We present the following article in accordance with the MDAR reporting checklist (available at https://atm.amegroups.com/article/view/10.21037/atm-22-5184/rc).
Methods
Screening for prognostic-related miRNAs
The miRNA expression data and clinical characteristics of cervical cancer [i.e., cervical squamous cell carcinoma (CESC) and endocervical adenocarcinoma] samples measured by Illumina-Hiseq were retrieved from TCGA database (https://portal.gdc.cancer.gov/), including 2,200 miRNAs obtained from 304 tumor samples. After normalizing the expression profile and removing the lowly expressed miRNAs (reads per million <1 in more than half of the tumor samples), the remaining miRNAs (n=459) underwent univariate Cox proportional hazard regression analyses, and miRNAs with P values evaluated by 1,000 permutations <0.05 (by the likelihood ratio test) were chosen for the subsequent analysis. A univariate Cox regression analysis was conducted to identify the tumor, node, metastasis (TNM) clinical characteristics that were risk factors correlated with the poor prognosis of CESC. Next, all 459 miRNAs were also used a multivariate Cox model adjusted according to the degree of TNM in the CESC patients. The significant miRNAs (P<0.05, n=56) were considered independent prognostic factors for CESC. We respectively selected the top 30 miRNAs in univariate and multivariate Cox regression analyses, the overlapping miRNAs (n=23) were considered to be prognostic-related miRNAs. The univariate and multivariate Cox regression analyses were performed using the R package ‘survival’ from CRAN. The Kaplan-Meier plots were generated by R package survminer. All the analyses were conducted with R software (version 4.1.0).
The discovery of miR-95-3p target genes in CESC
The potential target genes of miR-95-3p were predicted by the TargetScan (26), miRbase (27), and miRTarBase (28) databases. The CESC samples were divided into the following 3 groups according to the expression of miR-95-3p: (I) the miR-95-3p-high group (n=101); (II) the miR-95-3p-low group (n=101); and (III) the miR-95-3p-moderate group (n=102). A differential expression analysis was performed by DESeq2 (29) from the Bioconductor project of the miR-95-3p-high group and miR-95-3p-low groups. We chose VCAM1 (which overlapped in the miR-95-3p target genes and cell adhesion molecule-related genes and was correlated with a good prognosis in CESC patients) as the potential target genes of miR-95-3p in CESC.
GO and KEGG pathway enrichment analyses
The R package clusterProfiler (30) from the Bioconductor project was used to conduct the gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses. An adjusted P value <0.05 was considered statistically significant. A gene set enrichment analysis (GSEA) was performed by R “enrichplot” package from the Bioconductor project.
Cell culture and transfection
HeLa cells and SiHa cells were obtained from the Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences, Shanghai China and cultured in Dulbecco’s modified Eagle medium (Solarbio) containing 10% fetal bovine serum (BIOIND) at 37 ℃ in 5% carbon dioxide. The mimic (UUCAACGGGUAUUUAUUGAGCA) and the inhibitor (UGCUCAAUAAAUACCCGUUGAA) of miR-95-3p were purchased from Genebio (Shanghai, China). The miRNA mimic and inhibitor were transfected into the cells using LipofectamineTM 3000 in accordance with the manufacturer’s instructions. After transfecting for 12 h, the medium was changed and replaced with fresh cell culture medium before the subsequent experiments were performed.
RNA extraction and qRT-PCR
Total RNA was extracted by RNAiso plus Reagent (Thermo) and reverse transcribed into complementary deoxyribonucleic acid (DNA) by a reverse transcription kit (Servicebio). Quantitative real-time–polymerase chain reaction (qRT-PCR) was performed using 2× Synergy Brands Inc (SYBR) Green (Servicebio) on the Roche LightCycler480 II RT-PCR Detection System. The miRNAs were quantified by stem-loop RT-PCR, and U6 small nuclear RNA (U6) was used as an internal reference. All the reactions were triplicated and were calculated using the comparative threshold method (2−ΔΔCt). The sequences of the qRT-PCR primers are listed in Table S1.
CCK-8 cell viability assays
The HeLa and SiHa cells were transfected with either a miRNA inhibitor or mimic. After 48 h, the cells were evenly passaged to 96-well plates with 2×103 cells per well. The cells were cultured for 24, 48, 72 and 96 h, and 10 µL/well of Cell Counting Kit-8 (CCK-8) solution was added to the cells, which were then incubated for 1 h. The medium was then discarded, and cell proliferation was analyzed by measuring the absorption at 510 nm. The cell growth curves were depicted by Graphpad Prism 7 software.
Colony formation assays
The inhibitor and mimic of miR-95-3p were respectively transfected into the HeLa and SiHa cells. After 48 h, the cells were evenly plated into 12-well plates (200 cells per well). The cell culture medium was changed every 2 days. After 2 weeks, the cells were fixed with 4% paraformaldehyde for 15 min, then washed 3 times by phosphate buffered solution (PBS), and dyed with crystal violet staining solution for 30 min.
Wound-healing assays
The HeLa and SiHa cells were transfected with an inhibitor or mimic of miR-95-3p. After 48 h, the cells were dissociated by trypsin and re-seeded to 6-well plates with 10% serum culture medium. When the cells grew to 100% confluence, the cells were starved for 12 h and a straight wound was made by a micropipette tip in each well. Next, the floating cells and debris were washed by PBS for several times, and the cells were cultured with 1% serum culture medium to allow wound healing. Photographs were taken by microscopy at 0 and 48 h after wounding.
Transwell migration assays
The HeLa and SiHa cells were transfected with either an inhibitor or mimic of miR-95-3p for 48 h. Next, 200 µL of the cells in culture medium without serum were seeded in the upper compartment of Boyden chambers (6.5 mm pore size; Costar), and lower chambers were filled with 600 µL of the medium containing 20% fetal bovine serum. After 24 h of incubation, the upper side of the chamber was wiped with a cotton swab. The migratory cells on the bottom surface of the chamber were fixed with 4% paraformaldehyde for 25 min and stained with 0.1% crystal violet staining solution for 30 min. Finally, 3 randomly selected fields from each insert were photographed and counted under a light microscope (Olympus, Japan).
Western blot assays
The total protein in the cells was extracted. The protein was mixed with loading buffer and boiled for 5 min, and then treated with 10% sodium dodecylsulfate-polyacrylamide gel electrophoresis and transferred to the membranes. The membranes were blocked with 5% skimmed milk for 1 h, probed with primary antibodies VCAM1 (1:1,000, abcam), glyceraldehyde-3-phosphate dehydrogenase (GAPDH) (1:2,000, Sigma, St. Louis, MO, USA), and re-probed with a secondary antibody of either Alexa Fluor 680 or IRDye 800-conjugated secondary antibody. The gel was detected by the Odyssey system (LI-COR Biosciences, Lincoln, NE, USA). The protein images were analyzed by ImageJ2x software.
Immunohistochemical staining
For the immunohistochemical staining, the cervical cancer tissues were obtained from The Second People’s Hospital of Jiaozuo. The cervical cancer tissue samples was stained with an anti-VCAM1 antibody (1:100 dilution; Santa Cruz Biotechnology, Santa Cruz, CA, USA) and incubated for 1 h. The cells were the incubated for 30 min with horseradish-peroxidase-labeled goat-anti-rabbit secondary antibody (Invitrogen), and the section was colored using a DAB Horseradish Peroxidase Color Development (DAB) kit (Roche). VCAM1 protein expressions were imaged using a microscope (Olympus), and the data were reported automatically to a linked clinical database.
Ethical statement
The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The Institutional Review Board of The Second People’s Hospital of Jiaozuo approved this study (KY2021-04-047), and all enrolled patients who met the inclusion criteria signed an informed consent form.
Statistical analysis
All experiment results are presented as mean ± standard deviation (SD) from 3 independent experiments and showed successful reproducibility. All graphs were generated using GraphPad Prism7 (64-bit, La Jolla, CA, USA). Two-tailed unpaired t-tests (Student’s t-test) were used to obtain the P values. The data are presented as the mean ± standard deviation. *P<0.05, **P<0.01, ***P<0.001.
Results
MiR-95-3p was significantly correlated with the poor prognosis of CESC
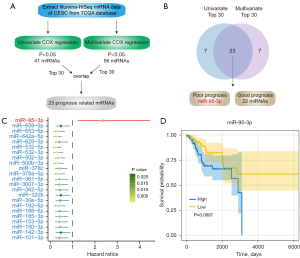
As Figure 1A shows, we downloaded the miRNA HiSeq data of CESC and endocervical adenocarcinoma from TCGA database and performed a univariate COX regression analysis to identify the clinical characteristics and miRNAs significantly correlated with CESC prognosis. We identified 41 prognosis-related miRNAs and also found that lymph node metastasis (N) and distal metastasis (M) were significantly correlated with a poor prognosis in CESC. Next, we included TNM status as a covariate in the multivariate Cox regression analysis, and 56 miRNAs that were significantly correlated with the prognosis of CESC were filtered out. We integrated the results of the two Cox regression analyses, ranked the miRNAs according to their P values, and selected the top 23 intersecting miRNAs (Figure 1B). Finally, we identified these miRNAs that were significantly correlated with CESC prognosis, of which 22 miRNAs were related to a good prognosis, and only 1 (i.e., miR-95-3p) was related to a poor prognosis (Figure 1C,1D). Collectively, these data showed that a high expression of miR-95-3p was correlated with a poor prognosis in CESC.
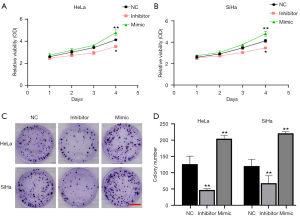
MiR-95-3p inhibition suppresses cell proliferation and migration in cervical cancer cell lines
There is emerging evidence that miR-95-3p plays vital roles in different cancer types (24-26). However, the roles of miR-95-3p in cervical cancer remain unknown. To further investigate the effects of miR-95-3p on the malignant behavior of cervical cancer cell, we first designed the inhibitor and mimic of miR-95-3p. The efficiencies of the inhibitor and mimic were confirmed by qRT-PCR in the HeLa and SiHa cells (Figure S1). The CCK-8 assays showed that the inhibition of miR-95-3p significantly suppressed cell proliferation in the HeLa and SiHa cells (Figure 2A,2B), and the cell colony assays further confirmed these results (Figure 2C,2D). The results indicated that miRNA-95-3p may play crucial roles in the malignant behavior of cervical cancer cells.
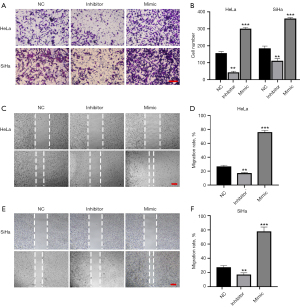
We also examined the role of miR-95-3p downregulation in cell migration. The transwell and wound-healing assays showed that miR-95-3p inhibition suppressed the cell migration ability of the HeLa and SiHa cells (Figure 3). As a result, the downregulation of miR-95-3p significantly decreased the cell viability of the 2 cervical cancer cell lines. To sum up, miR-95-3p acts as a critical factor in the occurrence and development of cervical cancer.
The overexpression of miR-95-3p induces the malignant behavior of cervical cancer cells
To explore whether the overexpression of miR-95-3p shows the opposite phenotype of miR-95-3p inhibition in cervical cancer cells, we further examined the effect of miR-95-3p upregulation on cell proliferation and cell migration in the HeLa and SiHa cells. Consistent with the above-mentioned results, the CCK-8 and colony formation assays showed that miR-95-3p overexpression significantly promoted the growth rate of both the HeLa and SiHa cells compared to that of the control cells transfected with the negative control (NC) (Figure 2A,2B). Further, the transwell and wound-healing assays also confirmed that miR-95-3p overexpression significantly promoted the cell movement of HeLa and SiHa cells (Figure 3). Collectively, we demonstrated that miR-95-3p is a poor prognosis-related marker of cervical cancer that promotes cell proliferation and migration.
VCAM1 is a target gene of miR-95-3p and is associated with good prognosis of cervical cancer
To investigate the underlying mechanisms of miR-95-3p in inducing malignant behavior in cervical cancer cells, we sorted the expression levels of miR-95-3p in TCGA database and divided them into 3 equal groups (i.e., the miR-95-3p-high, miR-95-3p-low, and miR-95-3p-moderate groups). Next, we performed a differential expression analysis of protein-coding genes in the miR-95-3p-high and miR-95-3p-low groups and identified 363 upregulated genes and 679 downregulated genes (|log2 fold change| >0.585, adjusted P value <0.05) (Figure 4A, Figure S2A).
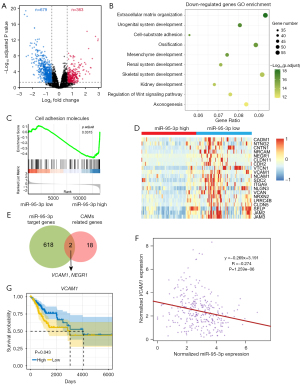
We then conducted a GO enrichment analysis. The GO terms among the upregulated genes included myeloid blood cell migration, epidermis development, and immune response (Figure S2B). The GO terms among the downregulated genes included extracellular matrix organization, cell-substrate adhesion and organ development (Figure 4B). Our previous data demonstrated that miR-95-3p promotes the migration of cervical cancer cells. Thus, we hypothesized that miR-95-3p promotes the migration of cancer cells by inhibiting the expression of cell adhesion-related genes.
We also found that the repressed KEGG pathways included cell adhesion molecules (Figure 4C). Further, 20 genes involved in cell adhesion molecules were downregulated in miR-95-3p-high expression patients (Figure 4D). To determine which of these cell adhesion molecule–related genes were downstream targets of miR-95-3p, we searched for the putative target genes of miR-95-3p in 3 online miRNA target predication databases (i.e., TargetScan, miRTarBase, and miRDB). These databases comprised a total of 620 target genes of miR-95-3p. We intersected these target genes with the cell adhesion molecule-related genes downregulated in patients with high miR-95-3p expression and found that VCAM1 and neuronal growth regulator 1 (NEGR1) were potential targets of miR-95-3p in cervical cancer cells (Figure 4E).
In addition, by accessing the cervical cancer samples in TCGA database, we found that the expression of VCAM1 and NEGR1 was negatively correlated with the expression of miR-95-3p (Figure 4F, Figure S2C). However, a high expression of VCAM1 indicated a good prognosis in cervical cancer patients, but high expression of NEGR1 was not associated with cervical cancer prognosis (Figure 4G, Figure S2D). In sum, these results suggested that VCAM1 is a target gene of miR-95-3p in cervical cancer, and miR-95-3p promotes the malignant behavior of cervical cancer cells by (at least in part) inhibiting the expression of VCAM1.
VCAM1 is negatively correlated with the expression of miR-95-3p, and its expression gradually decreases with tumor progression
VCAM1 is generally detected in endothelial cells, is activated by inflammatory factors (31), and mediates leukocyte-endothelial cell adhesion and signal transduction (32). The high expression of this membrane protein not only plays a key role in the development of some inflammatory diseases, such as atherosclerosis (33) and rheumatoid arthritis (34), but also exhibits an important role in the progression of human tumors (35-38).
Interestingly, in a previous analysis, we showed that the high expression of VCAM1 indicates a good prognosis in cervical cancer patients, which is in line with the results of a previous study (39). To examine the aberrant expression of VCAM1 in cervical tumor tissues and further explore the relationship between miR-95-3p and VCAM1, we collected clinical cervical tumor tissue samples and downregulated the expression of miR-95-3p with an inhibitor in the HeLa cells. The Western blot analysis showed that the knockdown of miR-95-3p in HeLa cells resulted in increased VCAM1 protein expression compared to the control cells (Figure 5A). The same result was confirmed by qRT-PCR in the HeLa cells (Figure 5B).
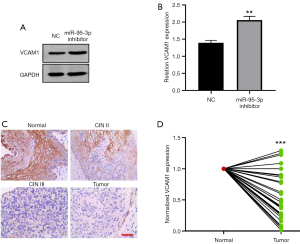
We also examined the expression of VCAM1 in cervical tumor tissue samples and matched adjacent normal cervical epithelial tissue samples using a immunohistochemical analysis and qRT-PCR. Representative immunohistochemical VCAM1 staining patterns are shown in Figure 5C. VCAM1-positive staining was predominantly localized to the cytoplasm in normal cells (Figure 5C). Intriguingly, the immunohistochemical analysis showed that the expression of VCAM1 in the Cervical Intraepithelial Neoplasia (CIN) III tissue sample was much lower than that in the CIN II tissue sample. Additionally, in the cervical tumor cells, VCAM1 showed barely any expression (Figure 5C). Similarly, the qRT-PCR results showed that VCAM1 was highly expressed in paired adjacent normal cervical epithelium tissue samples, but lowly expressed in cervical tumor tissue samples (Figure 5D). These results suggested that the expression of VCAM1 was negatively correlated with the expression of miR-95-3p, and the expression of VCAM1 was negatively correlated with tumor progression in cervical cancer patients.
Discussion
Cervical cancer is a common cancer in women worldwide and has a poor prognosis. The mechanism of carcinogenesis and development of cervical cancer remain unknown. In this study, we sought to identify any special molecules that could serve as prognostic markers of cervical cancer and to explore their potential biological functions and mechanisms.
The altered expression of miRNAs has been observed in various human diseases, including cancer. miRNAs participate in cell proliferation, invasion and metastasis, angiogenesis, and chemoresistance processes by regulating the target genes (6-8). Many studies have shown that the dysregulation of miRNAs is related to cervical cancer progression (40-42). Further, miRNAs have been confirmed to be more stable than messenger RNAs, thus miRNAs could be appropriate biomarkers for prognosis evaluations and have the therapeutic potential of cervical cancer (42). Tumor microenvironment (TME) plays a vital role in the growth and development of tumors, including immune cells, stromal cells, blood vessels, and extracellular matrix (ECM) (43). Immune cells and stromal cells dominate the tumor microenvironment in cervical cancer, and T cells and mast cells have been suggested to be significantly associated with the survival rate of patients with cervical cancer (44).
In the current study, by conducting comprehensive analyses of large-scale miRNA expression data and CESC clinical data from TCGA database, we successfully identified miR-95-3p as a candidate prognostic marker for cervical cancer. A high expression level of miR-95-3p indicated worse overall survival for CESC patients. A previous study showed that miR-95-3p was upregulated in prostatic cancer cells and was associated with the poor prognosis of prostatic cancer patients (45). Research has also shown that miR-95-3p promotes the development of prostatic cancer by targeting DKK3 and activating the Wnt/β-catenin pathway (45). Additionally, miR-95-3p has been reported to be an oncogene that promotes tumorigenesis by targeting p21 expression in hepatocellular carcinoma (46).
Similarly, miR-95-3p has also been shown to be associated with chemoresistance; researchers have hypothesized that cisplatin resistance develops via the downregulation of EMP1 and the increased phosphorylation of the PI3K/Akt pathway in gastric cancer (47). Conversely, miR-95-3p was found to inhibit cell proliferation and metastasis in colorectal carcinoma by targeting heparin binding growth factor (HDGF) (48). Thus, miR-95-3p plays bivalent roles in different cancer types. But the correlation between miR-95-3p and the tumor microenvironment is unclear. In the present study, we confirmed that the high expression level of miR-95-3p indicated a poor prognosis in CESC and promoted cervical cancer cell proliferation and migration. Future studies on the regulation of functional impact of miR-95-3p on apoptosis of cervical cancer cells and in vivo are clearly needed.
We also identified the cell adhesion molecule-related gene VCAM1 as the potential target of miR-95-3p. In previous studies, VCAM1 was shown to act as a vascular cell adhesion factor that helps regulate inflammation-associated vascular adhesion and the transendothelial migration of leukocytes, and it was conjected that it was an oncogene in other cancer types (36,49,50). And VCAM1 engagement regulates a critical immune-checkpoint in the bone marrow, and offers an alternative strategy to eliminate cancer cells via modulation of the innate immune tolerance (51). However, in our study, VCAM1 was associated with a good prognosis in cervical cancer, and the knockdown of miR-95-3p in the HeLa cells resulted in an increase in VCAM1 expression compared to the control. The results indicated that miR-95-3p promotes the malignant behavior of cervical cancer cells by inhibiting the expression of VCAM1. In subsequent research, we will add the dual luciferase reporter system to verify the targeted regulation of miR-95-3p to VCAM1.
Further, the qRT-PCR and immunohistochemical analysis results showed that VCAM1 was more lowly expressed in the cervical cancer tissues than the matched adjacent normal cervical epithelial tissues. However, the mechanism by which miR-95-3p promotes the malignant behavior of cervical cancer requires further investigation, and other downstream target genes may be involved in regulating the migration and invasion of cervical cancer cells.
In addition to the cell adhesion molecule pathway-related genes, the upregulated genes in patient samples with high miR-95-3p expression may also have target genes that regulate the malignant behavior of cervical cancer cells, and the specific role of VCAM1 in cervical cancer cells also requires further exploration. Also, miR-95-3p-VCAM1 axis may be regulated by lncRNAs, mRNAs and lncRNAs form endogenous competition by competitive binding to the same miRNA, and we will explore which lncRNAs are involved in this process in the future. Our findings nonetheless provide a novel perspective for the exploration of the prognostic biomarkers for cervical cancer and mechanisms underlying malignant behaviors of cervical cancer.
In summary, miR-95-3p affects the prognosis of cervical cancer patients by inhibiting the expression of the cell adhesion molecule-related gene VCAM1. Subsequent experiments place great emphasis on determining the functions of miR-95-3p in cervical cancer cell lines. However, further research needs to be conducted to discover the other biological functions of the downstream target genes of miR-95-3p. Overall, we provided the novel insight that miR-95-3p may serve as a prognostic biomarker for cervical cancer and could become a potential therapeutic target in treatment of cervical cancer.
Acknowledgments
Funding: This work was supported by the Fundamental Research Funds for the Universities of Henan Province (No. NSFRF210310), the Doctoral Fund of Henan Polytechnic University (No. B2020-51), the National Natural Science Foundation of China (No. 32100951), the Key Scientific Research Project for Higher Education of Henan Province of China (No. 23B310004), Medical Science and Technology Planning (Joint construction) Project of Henan Province (No. LHGJ20191350), and the Science and Technology Project of Henan Province of China (No. 212102310222).
Footnote
Reporting Checklist: The authors have completed the MDAR reporting checklist. Available at https://atm.amegroups.com/article/view/10.21037/atm-22-5184/rc
Data Sharing Statement: Available at https://atm.amegroups.com/article/view/10.21037/atm-22-5184/dss
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://atm.amegroups.com/article/view/10.21037/atm-22-5184/coif). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The Institutional Review Board of The Second People’s Hospital of Jiaozuo approved this study (KY2021-04-047), and all enrolled patients who met the inclusion criteria signed an informed consent form.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Cohen PA, Jhingran A, Oaknin A, et al. Cervical cancer. Lancet 2019;393:169-82. [Crossref] [PubMed]
- Hu Z, Ma D. The precision prevention and therapy of HPV-related cervical cancer: new concepts and clinical implications. Cancer Med 2018;7:5217-36. [Crossref] [PubMed]
- Park S, Kim J, Eom K, et al. microRNA-944 overexpression is a biomarker for poor prognosis of advanced cervical cancer. BMC Cancer 2019;19:419. [Crossref] [PubMed]
- Castellsagué X, Díaz M, de Sanjosé SInternational Agency for Research on Cancer Multicenter Cervical Cancer Study Group, et al. Worldwide human papillomavirus etiology of cervical adenocarcinoma and its cofactors: implications for screening and prevention. J Natl Cancer Inst 2006;98:303-15. [Crossref] [PubMed]
- Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell 2009;136:215-33. [Crossref] [PubMed]
- Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004;116:281-97. [Crossref] [PubMed]
- Singh G, Storey KB. MicroRNA Cues from Nature: A Roadmap to Decipher and Combat Challenges in Human Health and Disease? Cells 2021;10:3374. [Crossref] [PubMed]
- Shi Y, Zhao X, Hsieh J, et al. MicroRNA regulation of neural stem cells and neurogenesis. J Neurosci 2010;30:14931-6. [Crossref] [PubMed]
- Asadi-Moghaddam K, Chiocca EA, Lawler SE. Potential role of miRNAs and their inhibitors in glioma treatment. Expert Rev Anticancer Ther 2010;10:1753-62. [Crossref] [PubMed]
- Hill M, Tran N. miRNA interplay: mechanisms and consequences in cancer. Dis Model Mech 2021;14:dmm047662. [Crossref] [PubMed]
- Cheng L, Bao S, Rich JN. Potential therapeutic implications of cancer stem cells in glioblastoma. Biochem Pharmacol 2010;80:654-65. [Crossref] [PubMed]
- Nagamitsu Y, Nishi H, Sasaki T, et al. Profiling analysis of circulating microRNA expression in cervical cancer. Mol Clin Oncol 2016;5:189-94. [Crossref] [PubMed]
- Gao C, Zhou C, Zhuang J, et al. MicroRNA expression in cervical cancer: Novel diagnostic and prognostic biomarkers. J Cell Biochem 2018;119:7080-90. [Crossref] [PubMed]
- González-Quintana V, Palma-Berré L, Campos-Parra AD, et al. MicroRNAs are involved in cervical cancer development, progression, clinical outcome and improvement treatment response Oncol Rep 2016;35:3-12. (Review). [Crossref] [PubMed]
- Li J, Liu Q, Clark LH, et al. Deregulated miRNAs in human cervical cancer: functional importance and potential clinical use. Future Oncol 2017;13:743-53. [Crossref] [PubMed]
- Liu X, Zhou Y, Ning YE, et al. MiR-195-5p Inhibits Malignant Progression of Cervical Cancer by Targeting YAP1. Onco Targets Ther 2020;13:931-44. [Crossref] [PubMed]
- Chen X, Cao R, Liu H, et al. MicroRNA-15a-5p-targeting oncogene YAP1 inhibits cell viability and induces cell apoptosis in cervical cancer cells. Int J Mol Med 2020;46:1301-10. [Crossref] [PubMed]
- Wang A, Xu Q, Sha R, et al. MicroRNA-29a inhibits cell proliferation and arrests cell cycle by modulating p16 methylation in cervical cancer. Oncol Lett 2021;21:272. [Crossref] [PubMed]
- Lao G, Liu P, Wu Q, et al. Mir-155 promotes cervical cancer cell proliferation through suppression of its target gene LKB1. Tumour Biol 2014;35:11933-8. [Crossref] [PubMed]
- Liao H, Pan Y, Pan Y, et al. MicroRNA-874 is downregulated in cervical cancer and inhibits cancer progression by directly targeting ETS1. Oncol Rep 2018;40:2389-98. Retraction in: Oncol Rep 2022;48:165. [Crossref] [PubMed]
- Cline MS, Craft B, Swatloski T, et al. Exploring TCGA Pan-Cancer data at the UCSC Cancer Genomics Browser. Sci Rep 2013;3:2652. [Crossref] [PubMed]
- Zhang D, Duan Y, Cun J, et al. Identification of Prognostic Alternative Splicing Signature in Breast Carcinoma. Front Genet 2019;10:278. [Crossref] [PubMed]
- Lai J, Wang H, Pan Z, et al. A novel six-microRNA-based model to improve prognosis prediction of breast cancer. Aging (Albany NY) 2019;11:649-62. [Crossref] [PubMed]
- Bao C, Lu Y, Chen J, et al. Exploring specific prognostic biomarkers in triple-negative breast cancer. Cell Death Dis 2019;10:807. [Crossref] [PubMed]
- Bao C, Chen J, Chen D, et al. MiR-93 suppresses tumorigenesis and enhances chemosensitivity of breast cancer via dual targeting E2F1 and CCND1. Cell Death Dis 2020;11:618. [Crossref] [PubMed]
- Agarwal V, Bell GW, Nam JW, et al. Predicting effective microRNA target sites in mammalian mRNAs. Elife 2015;4:e05005. [Crossref] [PubMed]
- Kozomara A, Birgaoanu M, Griffiths-Jones S. miRBase: from microRNA sequences to function. Nucleic Acids Res 2019;47:D155-62. [Crossref] [PubMed]
- Hsu SD, Lin FM, Wu WY, et al. miRTarBase: a database curates experimentally validated microRNA-target interactions. Nucleic Acids Res 2011;39:D163-9. [Crossref] [PubMed]
- Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 2014;15:550. [Crossref] [PubMed]
- Yu G, Wang LG, Han Y, et al. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 2012;16:284-7. [Crossref] [PubMed]
- Vonderheide RH, Tedder TF, Springer TA, et al. Residues within a conserved amino acid motif of domains 1 and 4 of VCAM-1 are required for binding to VLA-4. J Cell Biol 1994;125:215-22. [Crossref] [PubMed]
- Yamada Y, Arao T, Matsumoto K, et al. Plasma concentrations of VCAM-1 and PAI-1: a predictive biomarker for post-operative recurrence in colorectal cancer. Cancer Sci 2010;101:1886-90. [Crossref] [PubMed]
- Galkina E, Ley K. Vascular adhesion molecules in atherosclerosis. Arterioscler Thromb Vasc Biol 2007;27:2292-301. [Crossref] [PubMed]
- Carter RA, Wicks IP. Vascular cell adhesion molecule 1 (CD106): a multifaceted regulator of joint inflammation. Arthritis Rheum 2001;44:985-94. [Crossref] [PubMed]
- Yurkovetsky Z, Skates S, Lomakin A, et al. Development of a multimarker assay for early detection of ovarian cancer. J Clin Oncol 2010;28:2159-66. [Crossref] [PubMed]
- Zhang D, Bi J, Liang Q, et al. VCAM1 Promotes Tumor Cell Invasion and Metastasis by Inducing EMT and Transendothelial Migration in Colorectal Cancer. Front Oncol 2020;10:1066. [Crossref] [PubMed]
- Chen Q, Zhang XH, Massagué J. Macrophage binding to receptor VCAM-1 transmits survival signals in breast cancer cells that invade the lungs. Cancer Cell 2011;20:538-49. [Crossref] [PubMed]
- Horn L, Dahlberg SE, Sandler AB, et al. Phase II study of cisplatin plus etoposide and bevacizumab for previously untreated, extensive-stage small-cell lung cancer: Eastern Cooperative Oncology Group Study E3501. J Clin Oncol 2009;27:6006-11. [Crossref] [PubMed]
- Qin R, Cao L, Ye C, et al. A novel prognostic prediction model based on seven immune-related RNAs for predicting overall survival of patients in early cervical squamous cell carcinoma. BMC Med Genomics 2021;14:49. [Crossref] [PubMed]
- Pardini B, De Maria D, Francavilla A, et al. MicroRNAs as markers of progression in cervical cancer: a systematic review. BMC Cancer 2018;18:696. [Crossref] [PubMed]
- He Y, Lin J, Ding Y, et al. A systematic study on dysregulated microRNAs in cervical cancer development. Int J Cancer 2016;138:1312-27. [Crossref] [PubMed]
- Mitra T, Elangovan S. Cervical cancer development, chemoresistance, and therapy: a snapshot of involvement of microRNA. Mol Cell Biochem 2021;476:4363-85. [Crossref] [PubMed]
- Anderson NM, Simon MC. The tumor microenvironment. Curr Biol 2020;30:R921-5. [Crossref] [PubMed]
- Zhang W, Xiao P, Tang J, et al. m6A Regulator-Mediated Tumour Infiltration and Methylation Modification in Cervical Cancer Microenvironment. Front Immunol 2022;13:888650. [Crossref] [PubMed]
- Xi M, Cheng L, Hua W, et al. MicroRNA-95-3p promoted the development of prostatic cancer via regulating DKK3 and activating Wnt/β-catenin pathway. Eur Rev Med Pharmacol Sci 2019;23:1002-11. [PubMed]
- Ye J, Yao Y, Song Q, et al. Up-regulation of miR-95-3p in hepatocellular carcinoma promotes tumorigenesis by targeting p21 expression. Sci Rep 2016;6:34034. [Crossref] [PubMed]
- Ni Q, Zhang Y, Tao R, et al. MicroRNA-95-3p serves as a contributor to cisplatin resistance in human gastric cancer cells by targeting EMP1/PI3K/AKT signaling. Aging (Albany NY) 2021;13:8665-87. [Crossref] [PubMed]
- Hong YG, Huang ZP, Liu QZ, et al. MicroRNA-95-3p inhibits cell proliferation and metastasis in colorectal carcinoma by HDGF. Biomed J 2020;43:163-73. [Crossref] [PubMed]
- Shen J, Zhai J, You Q, et al. Cancer-associated fibroblasts-derived VCAM1 induced by H. pylori infection facilitates tumor invasion in gastric cancer. Oncogene 2020;39:2961-74. [Crossref] [PubMed]
- Kong DH, Kim YK, Kim MR, et al. Emerging Roles of Vascular Cell Adhesion Molecule-1 (VCAM-1) in Immunological Disorders and Cancer. Int J Mol Sci 2018;19:1057. [Crossref] [PubMed]
- Pinho S, Wei Q, Maryanovich M, et al. VCAM1 confers innate immune tolerance on haematopoietic and leukaemic stem cells. Nat Cell Biol 2022;24:290-8. [Crossref] [PubMed]
(English Language Editor: L. Huleatt)


