
Upregulated miR-155 inhibits inflammatory response induced by C. albicans in human monocytes derived dendritic cells via targeting p65 and BCL-10
Introduction
Candida albicans (C. albicans) is an opportunistic fungal pathogen commonly colonizing on the surface of skin and mucosa (oral cavity, intestinal tract and vagina). It rarely causes infection in immunocompetent individuals (1); however, in immunocompromised individuals, C. albicans is one of the most prevalent fungal pathogen. It causes various types of candidiasis, ranging from superficial infection to invasive systemic infection (2). In recent years, with the increasing number of immunocompromised patients [e.g., cancer therapy, transplantation and the human immunodeficiency virus (HIV) epidemics], the incidence of C. albicans infections has considerably increased (3). In addition, physical alterations of the anatomical barrier, such as surgery, indwelling medical devices or antibiotic treatment, facilitate the access of fungi to the bloodstream and lead to increased incidence of disseminated candidiasis (4). Noteworthy, the nosocomial bloodstream infection in immunosuppressive patients is life-threatening, with a mortality rate as high as 40% (5).
The innate immune response is the first defense line against C. albicans infection (6). It is initiated by the recognition of the conserved molecular components of C. albicans, termed as pathogen-associated molecular patterns (PAMPs). The recognition of PAMPs depends on the pattern recognition receptors (PRRs) on innate immune cells (7). Dendritic cells (DCs), a kind of classical innate immune cells, plays an important role in anti-C. albicans immune response. With PRRs expressed on its surface, DCs senses the PAMPs of C. albicans and initiates the innate immune response rapidly. Many PRRs are involved in the interaction of C. albicans, such as toll-like receptors (TLRs), C-type lectin receptors (CLRs) and nucleotide-binding oligomerization domain (NOD)-like receptors (NLRs) (8). Dectin-1, a member of CLRs, can sense the β-glucan on the surface of C. albicans. The interaction between Dectin-1 and β-glucan triggers two intracellular signaling pathways named spleen tyrosine kinase (Syk)-dependent pathway and Raf-1 dependent pathway. The activation of these two pathways induces the immune response against C. albicans, which is characterized by the release of several types of pro-inflammatory cytokines, such as interleukin-6 (IL-6) and tumor necrosis factor-α (TNF-α) to control C. albicans infection (9,10).
MicroRNA (miRNA) is a type of short non-coding RNA that regulates multiple target genes expression at post-transcriptional level (11). It is widely involved in the regulation of immune cell development and activation (12,13). MiR-155 is an active immune regulator in innate immune response with dose-dependent effects (14,15). Moderate elevated miR-155 expression is critical for pro-inflammatory response; however, extremely high level of miR-155 starts to gradually terminate the response, due to its ability to target different transcripts (15). Previous studies have shown that miR-155 was upregulated by heat-killed C. albicans in murine bone marrow-derived macrophages (BMDMs) (16) and human DCs (17). However, the biological function of miR-155 in human monocytes derived DCs treated with C. albicans remains unknown. Hence, the aim of the present study was to investigate the biological function and underlying mechanisms of miR-155 in C. albicans activated DCs.
Methods
Candida albicans (C. albicans)
C. albicans (SC5314) strain was grown in Sabouraud dextrose (SBD) agar plates at 37 °C. For cells preparation, C. albicans cultures were incubated in Sabouraud broth at 37 °C overnight with shaking. Cells were harvested by centrifugation, washed twice with phosphate-buffered saline (PBS), and heat-killed for 1 hour at 100 °C (18). C. albicans were counted and adjusted to the proper concentration before using.
Cell lines and culture
Healthy individuals’ peripheral blood mononuclear cells (PBMCs) were isolated using density centrifugation with Ficoll (GE Healthcare, Sweden). CD14+ monocyte subsets were positively selected using anti-CD14 MACS microbeads from freshly isolated PBMCs, according to the instructions of the manufacturer (Miltenyi Biotec, Germany). Purified monocytes were cultured at 37 °C in 6-well plates with complete RPMI 1640 medium at the concentration of 106/mL. The cells were induced into immature DCs with 50 ng/mL of human granulocyte-macrophage colony-stimulating factor (GM-CSF, R&D Systems, USA) and 50 ng/mL human IL-4 (R&D Systems, USA) for 6 days and the culture was replaced at the third and fifth day of induction (19). This study was approved by the Ethic Board of Changzheng Hospital and written approvals were obtained from the subjects.
THP-1 cells, 293T cells and murine RAW264.7 cells were obtained from the Shanghai Institutes for Biological Sciences. All cells were cultured in either RPMI 1640 medium or DMEM (Gibco, USA) containing 10% fetal bovine serum (FBS, Gibco, USA) and 100 U/mL penicillin-streptomycin (Mediatech, USA).
Immature DCs were treated with heat-killed C. albicans at a ratio of 1:1 (20) or 100 ng/mL lipopolysaccharide (LPS, Sigma, USA). For inhibiting signaling pathways, DCs were pretreated with following inhibitors for 1 hour: Dectin-1 inhibitor laminarin 100 µg/mL, Syk inhibitor R406 5 µM, Raf-1 inhibitor GW5074 10 µM, MEK/ERK inhibitor U0126 10 µM, JNK inhibitor SP600125 10 µM and p38 kinase inhibitor SB203580 1 µM. All these inhibitors, except laminarin, were purchased from MedChem Express (USA), while laminarin was purchased from Sigma-Aldrich (USA).
Transient transfection
Small interfering RNAs (siRNAs) for Dectin-1, NF-κB p65 and B cell leukemia/lymphoma 10 (BCL-10) were ordered from RiboBio Company (Guangzhou, China). The sequences of siRNAs were presented in Table S1. MiR-155 mimics and mimic control, miR-155 inhibitor and inhibitor control were purchased from RiboBio Company, and used at the concentration of 50 and 100 nM, respectively. SiRNAs (100 nM) were transfected into DCs using riboFECTTM CP reagent from RiboBio according to the manufacturer’s instructions. Twenty-four or forty-eight hours after transfection, cells and the supernatants were harvested for the subsequent experiments. The transfected miRNA negative control (NC) tagged by red fluorescence was imaged by fluorescence microscopy.

Full table
RNA isolation and quantitative reverse-transcriptase polymerase chain reaction (qRT-PCR)
Total RNA, including miRNA, was extracted using Trizol reagent (Invitrogen, USA) according to the manufacturer’s instructions. RNA concentration was assessed by NanoDrop 2000 (Wilmington, DE, USA). Total RNA was reverse-transcribed to complementary DNA (cDNA) with PrimeScript™ RT Master Mix (Takara, Japan). qRT-PCR for target genes was performed on the Applied Biosystems 7500 Sequence (Applied Biosystems, USA). The primers were presented in Table S2. The primers for miR-155 and U6 small nuclear RNA were purchased from RiboBio Company (Guangzhou, China). Relative expression of messenger RNA (mRNA) and miRNA was calculated using the 2–ΔΔCT method (21).

Full table
Western blot
After stimulation and transfection, cells were harvested and centrifuged. Protein was isolated by RIPA Lysis Buffer containing protease inhibitor cocktail and phosphatase inhibitor. Protein concentration was quantified using BCA protein assay kit (Takara, Japan). Equal amounts of protein were separated by 10% SDS-PAGE (Beyotime Biotechnology) and transferred onto PVDF membranes (Millipore, USA). After blocking, the membranes were sequentially incubated with specific primary antibodies for 2 hours at room temperature. Antibodies against phospho-ERK (ab201015), total-ERK (ab184699), phospho-JNK (ab124956), total-JNK (ab179461) and BCL-10 (ab108328) were purchased from Abcam (USA); while phospho-p65 (3033S), Dectin-1 (60128S), p38 (9212S), phospho-p38 (4511S), c-Jun (9165S) and phospho-c-Jun (3270S) were from Cell Signaling Technology (USA); and total p65 [5006] was from Affinity Bioscience (USA). Immune-complexes were incubated with fluorescein-conjugated secondary antibodies and then detected using an Odyssey fluorescence scanner (Li-Cor, USA).
Enzyme-linked immunosorbent assay (ELISA)
The concentrations of cytokines in cell culture supernatants were measured using specific commercial ELISA kits (Neobioscience, China) according to the manufacturer’s instructions. All experiments were performed in triplicate.
Luciferase reporter assay
Luciferase reporter plasmids p65-3'UTR-Wt, p65-3'UTR-Mt, BCL-10-3'UTR-Wt, BCL-10-3'UTR-Mt were constructed from Obio Technology. Then these plasmids (0.2 µg) were co-transfected into 293T cells (2×105/mL) with a renilla control plasmid (0.01 µg) and miR-155 mimics (100 nM), NC (100 nM) using 0.2 µL Lipo2000 transfection reagent (Invitrogen, USA) according to the manufacture’s instruction. Medium contains transfection reagents was replaced with fresh and complete DMEM medium 24 hours after transfection. Reporter luciferase activities were measured using the Dual-Luciferase kit (E1910, Promega) according to the manufacture’s instruction 48 hours later.
Statistical analysis
All experiments were performed in triplicate. All statistical analyses were performed using SPSS 19.0 (SPSS Inc., Chicago, IL, USA) or GraphPad Prism 6.0 (GraphPad Software, La Jolla, CA, USA). Data were presented as mean and standard deviation (SD) from at least three independent experiments. Student’s t-test was used to compare the mean values between two groups. Multiple comparisons were performed using one-way analysis of variance (ANOVA). A P value <0.05 was considered statistically significant.
Results
Heat-killed C. albicans promoted the expression of pro-inflammatory factors and miR-155 in DCs
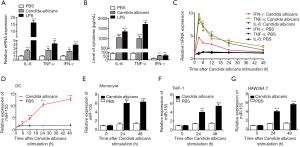
Heat-killed C. albicans significantly increased the expression of intracellular IL-6, TNF-α and interferon-γ (IFN-γ) mRNA as well as the concentration of these cytokines in culture media after 24 hours’ stimulation (Figure 1A,B). We further detected the dynamic changes of these cytokine’s mRNA with PCR, then found that their expressions were upregulated up to 3 hours after C. albicans stimulation and then decreased (Figure 1C). Meanwhile, miR-155 was significantly upregulated in DCs by heat-killed C. albicans in time dependent manner (Figure 1D). In addition, a significant increase of miR-155 expression was detected in human monocytes (Figure 1E), THP-1 cells (Figure 1F) and murine RAW264.7 cells (Figure 1G) in response to C. albicans for 24 and 48 hours.
Increased miR-155 expression in DCs induced by C. albicans was mediated via Dectin-1-Syk/Raf-1-MAPK pathway
To explore whether the upregulation of miR-155 was relevant to Dectin-1, DCs were treated with Dectin-1 blocker laminarin for 1 hour before heat-killed C. albicans stimulation. As shown in Figure 2A, the upregulation of miR-155 was significantly impaired by laminarin. Further, we specifically suppressed the expression of Dectin-1 in DCs using siRNA. Dectin-1 expression was significantly decreased by siRNA at mRNA and protein level (Figure S1A,B). The upregulation of miR-155 induced by C. albicans was significantly impaired by siRNA of Dectin-1 (Figure 2B). A previous study has indicated that the interaction of C. albicans and Dectin-1 could further activate Syk-dependent signaling pathway and Raf-1-dependent pathway (22). To explore whether these two pathways were involved in the upregulation of miR-155 upon Dectin-1 stimulation, DCs were pretreated with a specific inhibitor for Syk (R406) and Raf-1 (GW5074), respectively. As shown in Figure 2C, both R406 and GW5074 significantly attenuated the upregulation of miR-155 induced by C. albicans. These results demonstrate that the increased miR-155 expression in DCs induced by C. albicans is partially mediated via Dectin-1-Syk/Raf-1 pathway.
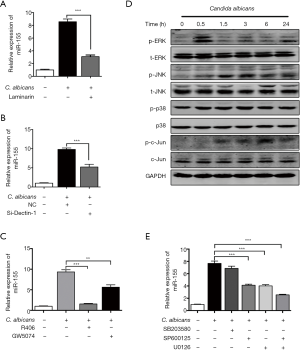
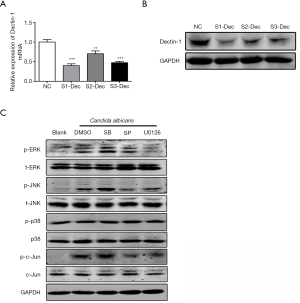
Next, we tested whether the induction of miR-155 by C. albicans was dependent on MAPK pathway using inhibitors. First, we found that both ERK and JNK, but not p38, in MAPK pathway were significantly phosphorylated in response to C. albicans (Figure 2D). We used the U0126, SP600125 and SB203580 to in inhibit the phosphorylation of MEK/ERK, JNK and p38, respectively (Figure S1C). The MEK/ERK inhibitor U0126 and the JNK inhibitor SP600125 significantly suppressed the upregulation of miR-155 by C. albicans stimulation, whereas the p38 inhibitor SB203580 had little influence on the expression of miR-155 (Figure 2E). When both ERK and JNK were inhibited, the expression of miR-155 was further declined (Figure 2E). These results imply that MAPK pathway is involved in the induction of miR-155 upon C. albicans stimulation.
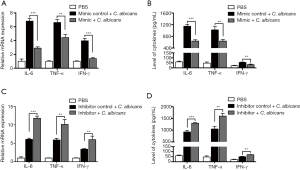
MiR-155 suppressed the upregulation of pro-inflammatory cytokines in DCs induced by heat-killed C. albicans
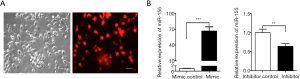
To investigate the function of miR-155 on the production of inflammatory cytokines, synthetic miR-155 mimics or scrambled control oligonucleotides were transfected into DCs. Transfection efficiency was detected by fluorescence microscope (Figure S2A) and qRT-PCR (Figure S2B) after 24 hours. Overexpression of miR-155 significantly suppressed C. albicans-induced expression and secretion of IL-6, TNF-α and IFN-γ (Figure 3A,B); whereas inhibition of endogenous miR-155 increased the expression and concentration of IL-6, TNF-α and IFN-γ induced by C. albicans (Figure 3C,D). These results indicate that the upregulation of miR-155 has a negative effect on the production of pro-inflammatory cytokines in heat-killed C. albicans treatment DCs.
MiR-155 downregulated the expression of IL-6, TNF-α and IFN-γ in DCs against C. albicans through targeting NF-κB p65 and BCL-10
To identify underlying mechanisms of miR-155 in inflammatory response induced by C. albicans, three types of bioinformatic prediction software, named Targetscan, miRanda and miRNA.org, were used to predict the potential targets of miR-155. Bioinformatics analysis showed that 3'UTR of NF-κB p65 contained miR-155 binding sites at 197–204 nucleotides (Figure 4A). Luciferase activity was significantly reduced when co-transfected with wild type of p65 and miR-155 (Figure 4A), indicating that miR-155 can bind to the 3'UTR of p65 mRNA directly. Then, upon C. albicans stimulation, miR-155 mimics reduced the protein level of p65, while miR-155 inhibitor had an opposite effect (Figure 4B). The dynamic changes of p65 in C. albicans treated DCs were also investigated. We found that the expression of p65 was upregulated within 6 hours after C. albicans stimulation but then decreased (Figure S3A). Furthermore, we used siRNA to knock down the expression of p65 and determined the production of IL-6, TNF-α and IFN-γ induced by C. albicans. We found that p65 siRNA significantly inhibit the expression of p65 at both mRNA and protein level (Figure S3B,C). The production of IL-6, TNF-α and IFN-γ was impaired by p65 siRNA in C. albicans treated DCs, at both mRNA (Figure 4C) and protein levels (Figure 4D). These results indicate miR-155 inhibits the production of IL-6, TNF-α and IFN-γ in C. albicans treated DCs partially by targeting p65.
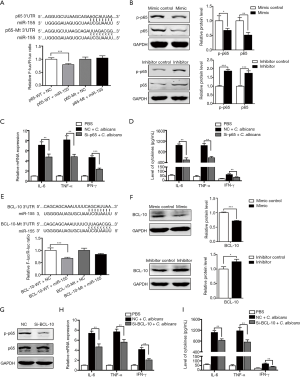
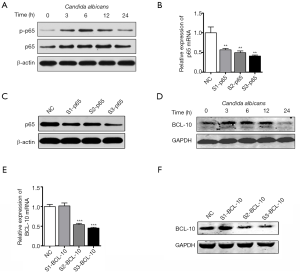
We found that the 3'UTR of BCL-10 contained miR-155 binding sites at 405–412 nucleotides (Figure 4E). Then luciferase activity was significantly reduced when co-transfected with BCL-10 wild type and miR-155 (Figure 4E). Upon C. albicans stimulation, protein level of BCL-10 was reduced after transfection with miR-155 mimics but increased with miR-155 inhibitor (Figure 4F). We also determined the dynamic changes of BCL-10 in C. albicans treated DCs. Similar to p65, the expression of BCL-10 increased up to 6 hours after C. albicans stimulation but decreased then (Figure S3D). We used siRNA to knock down the expression of BCL-10 and found that its expression was inhibited by siRNA (Figure S3E,F). Notably, siRNA of BCL-10 decreased the phosphorylation of p65 (Figure 4G). The levels of IL-6, TNF-α and IFN-γ induced by C. albicans were repressed significantly by siRNA of BCL-10 (Figure 4H,I). Taken together, these results indicate that miR-155 inhibits phosphorylation of p65 and inflammatory cytokines secretion by targeting BCL-10.
Discussion
In this study, we found that heat-killed C. albicans could upregulate the expression of miR-155 and inflammatory cytokines in DCs. Up-regulated miR-155 negatively regulated the production of inflammatory cytokines by targeting p65 and BCL-10. These results indicate that miR-155 plays a crucial role in negatively regulating the inflammation response against C. albicans in DCs.
Some previous studies have detected the miRNA profile of C. albicans treated cells. Using miRNA array and qRT-PCR, Monk et al. (16) reported that the expression of miR-155 was increased in murine BMDMs stimulated with C. albicans, but the function of elevated miR-155 was not illustrated. Similar to the study performed by Dix et al. (17), we observed the upregulation of miR-155 in human DCs treated with C. albicans. Besides, we found that heat-killed C. albicans could also increase miR-155 in human monocytes, THP-1 cells and RAW 264.7 cells. Interestingly, previous studies reported that miR-155 could also be elevated by viral infection or LPS in monocytes, macrophages and DCs (23-28). Taken together, these findings revealed that miR-155 may be upregulated in innate immune cells by various pathogens.
Dectin-1 was confirmed to bind β-glucans on C. albicans wall, playing an important role in the phagocytosis of fungi (29) and activation of immune response (30). In our study, the expression of miR-155 was attenuated by Dectin-1 inhibitor laminarin and siRNA of Dectin-1, indicating that the induction of miR-155 by C. albicans is partially through the activation of Dectin-1. A previous study has shown that the increased miR-155 in murine macrophages exposure to UV-killed C. albicans hyphae was associated with Dectin-1-Syk pathway, while Raf-1 pathway had no effect on miR-155 expression (31). In contrast, using specific inhibitors for each pathway, our study demonstrates that both of Dectin-1-Syk pathway and Dectin-1-Raf-1 pathway are involved in the induction of miR-155. The inconsistence between our work and previous study might be attributed to the different strains of C. albicans, inactivated methods and cell lines used in experiments.
MAPK signal pathways are known to be triggered by inflammatory response induced by Dectin-1 (32,33). Notably, MAPK pathways were reported to modulate miR-155 expression in murine macrophages by poly-I:C or TNF-α stimulation (34). In addition, the promoter regions of miR-155 primary transcript, B-cell integration cluster (BIC) contain binding sites for activator protein-1 (AP-1) (35). In this study, ERK and JNK pathway were significantly phosphorylated when exposure to C. albicans. Inhibition of ERK or JNK significantly attenuated the enhanced expression of miR-155 stimulated by C. albicans, whereas p38 inhibitor had no effect on miR-155 accumulation. These might be attributed to that inhibition of ERK or JNK pathway could suppress the phosphorylation of c-Jun (AP-1 key component), influencing the transcription of miR-155. Since cross-talk is existed among MAPK pathways, inhibition of one pathway could activate another (36). Our study found that both ERK and JNK were inhibited, the expression of miR-155 was further declined. Previous studies have reported that miR-155 could be induced by inflammatory cytokines, such as TNF-α or IFN-β (34,37,38). Therefore, we hypothesize that there might be two possible mechanisms responsible for the increased expression of miR-155. One is the direct activation of Dectin-1 and its downstream intracellular signaling pathways by C. albicans; another is the effect of inflammatory cytokines released by innate immune cells.
As miR-155 has been reported to be involved in regulating the strength and duration of innate immune response (14,39), whether miR-155 could regulate the strength of innate immune response triggered by C. albicans in DCs remained unknown. We found that miR-155 suppressed the production of inflammatory cytokines induced by C. albicans, including IL-6, TNF-α and IFN-γ, indicating that miR-155 plays as a negative regulator in inflammation to C. albicans. To the best of our knowledge, this is the first study reporting that miR-155 negatively regulate the releasing of IFN-γ in DCs. Then we identified the underlying mechanisms of miR-155 in C. albicans induced inflammatory response. Previous studies have indicated that p65, a subunit of NF-κB, was the target of miR-155 in endothelial inflammation (40) and gastric cancer (39,41). Therefore, we hypothesize that miR-155 impairs the upregulation of inflammatory cytokines via targeting p65. Using reporter assay and western blot, we found that miR-155 repressed the expression of p65 by direct targeting its 3’UTR. Furthermore, knockdown of p65 reproduced the effect of miR-155 mimics. These results indicate that miR-155 negatively regulates inflammatory cytokines producing via targeting p65.
Previous studies have indicated that BCL-10 is involved in the immune response against C. albicans. After the Dectin-1 activation by C. albicans and phosphorylation by Syk, caspase recruitment domain-containing protein 9 (CARD9) forms a complex with BCL-10 and mucosa-associated lymphoid tissue 1 (MALT1), mediating NF-κB activation and initiating the expression of several type of inflammatory cytokines (42). Furthermore, bioinformatics analysis indicated that BCL-10 was a potential target of miR-155. Therefore, we hypothesize that BCL-10 mediates the anti-inflammation effect of miR-155. In this study, we found that miR-155 repressed the expression of BCL-10 induced by C. albicans, and BCL-10 was a direct target of miR-155. Furthermore, silencing BCL-10 by siRNA suppressed the phosphorylation of p65 and the secretion of pro-inflammatory cytokines, reproducing the anti-inflammation effects of miR-155. Accordingly, we deduce that miR-155 negatively regulated pro-inflammatory cytokines in response to C. albicans by functionally targeting BCL-10.
Conclusively, our study demonstrated that the activation of Dectin-1 pathway in DCs upregulated the expression of miR-155. With targeting NF-κB p65 and BCL-10, increased expression of miR-155 attenuated the excessive inflammation induced by C. albicans. MiR-155 might “fine-tune” the innate immune response triggered by C. albicans.
Acknowledgments
Funding: This work was supported by a grant from the National Basic Research Program of China (973 Program) (Grant No. 2013CB531603), and the National Natural Science Foundation of China (Grant Nos. 81302541, 81471608).
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
Ethical Statement: The authors are responsible for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. This study was approved by the Ethic Board of Changzheng Hospital and written approvals were obtained from the subjects.
References
- Kim J, Sudbery P. Candida albicans, a major human fungal pathogen. J Microbiol 2011;49:171-7. [Crossref] [PubMed]
- Pfaller MA, Diekema DJ. Epidemiology of invasive mycoses in North America. Crit Rev Microbiol 2010;36:1-53. [Crossref] [PubMed]
- Yapar N. Epidemiology and risk factors for invasive candidiasis. Ther Clin Risk Manag 2014;10:95-105. [Crossref] [PubMed]
- Spellberg B. Novel insights into disseminated candidiasis: pathogenesis research and clinical experience converge. PLoS Pathog 2008;4:e38. [Crossref] [PubMed]
- Khader SA, Gaffen SL, Kolls JK. Th17 cells at the crossroads of innate and adaptive immunity against infectious diseases at the mucosa. Mucosal Immunol 2009;2:403-11. [Crossref] [PubMed]
- Cheng SC, Joosten LA, Kullberg BJ, et al. Interplay between Candida albicans and the mammalian innate host defense. Infect Immun 2012;80:1304-13. [Crossref] [PubMed]
- Becker KL, Ifrim DC, Quintin J, et al. Antifungal innate immunity: recognition and inflammatory networks. Semin Immunopathol 2015;37:107-16. [Crossref] [PubMed]
- Zheng NX, Wang Y, Hu DD, et al. The role of pattern recognition receptors in the innate recognition of Candida albicans. Virulence 2015;6:347-61. [Crossref] [PubMed]
- Brown GD. Dectin-1: a signalling non-TLR pattern-recognition receptor. Nat Rev Immunol 2006;6:33-43. [Crossref] [PubMed]
- Gow NA, Netea MG, Munro CA, et al. Immune recognition of Candida albicans beta-glucan by dectin-1. J Infect Dis 2007;196:1565-71. [Crossref] [PubMed]
- Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004;116:281-97. [Crossref] [PubMed]
- O'Connell RM, Rao DS, Chaudhuri AA, et al. Physiological and pathological roles for microRNAs in the immune system. Nat Rev Immunol 2010;10:111-22. [Crossref] [PubMed]
- Nahid MA, Satoh M, Chan EK. MicroRNA in TLR signaling and endotoxin tolerance. Cell Mol Immunol 2011;8:388-403. [Crossref] [PubMed]
- Bandyopadhyay S, Long ME, Allen LA. Differential expression of microRNAs in Francisella tularensis-infected human macrophages: miR-155-dependent downregulation of MyD88 inhibits the inflammatory response. PLoS One 2014;9:e109525. [Crossref] [PubMed]
- Michaille JJ, Awad H, Fortman EC, et al. miR-155 expression in antitumor immunity: The higher the better? Genes Chromosomes Cancer 2019;58:208-18. [Crossref] [PubMed]
- Monk CE, Hutvagner G, Arthur JS. Regulation of miRNA transcription in macrophages in response to Candida albicans. PLoS ONE 2010;5:e13669. [Crossref] [PubMed]
- Dix A, Czakai K, Leonhardt I, et al. Specific and Novel microRNAs Are Regulated as Response to Fungal Infection in Human Dendritic Cells. Front Microbiol 2017;8:270. [Crossref] [PubMed]
- Smeekens SP, Gresnigt MS, Becker KL, et al. An anti-inflammatory property of Candida albicans beta-glucan: Induction of high levels of interleukin-1 receptor antagonist via a Dectin-1/CR3 independent mechanism. Cytokine 2015;71:215-22. [Crossref] [PubMed]
- Lu C, Huang X, Zhang X, et al. miR-221 and miR-155 regulate human dendritic cell development, apoptosis, and IL-12 production through targeting of p27kip1, KPC1, and SOCS-1. Blood 2011;117:4293-303. [Crossref] [PubMed]
- Toth A, Csonka K, Jacobs C, et al. Candida albicans and Candida parapsilosis induce different T-cell responses in human peripheral blood mononuclear cells. J Infect Dis 2013;208:690-8. [Crossref] [PubMed]
- Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001;25:402-8. [Crossref] [PubMed]
- Gringhuis SI, den Dunnen J, Litjens M, et al. Dectin-1 directs T helper cell differentiation by controlling noncanonical NF-kappaB activation through Raf-1 and Syk. Nat Immunol 2009;10:203-13. [Crossref] [PubMed]
- Tili E, Michaille JJ, Cimino A, et al. Modulation of miR-155 and miR-125b levels following lipopolysaccharide/TNF-alpha stimulation and their possible roles in regulating the response to endotoxin shock. J Immunol 2007;179:5082-9. [Crossref] [PubMed]
- Lu F, Weidmer A, Liu CG, et al. Epstein-Barr virus-induced miR-155 attenuates NF-kappaB signaling and stabilizes latent virus persistence. J Virol 2008;82:10436-43. [Crossref] [PubMed]
- O'Connell RM, Rao DS, Chaudhuri AA, et al. Sustained expression of microRNA-155 in hematopoietic stem cells causes a myeloproliferative disorder. J Exp Med 2008;205:585-94. [Crossref] [PubMed]
- Ceppi M, Pereira PM, Dunand-Sauthier I, et al. MicroRNA-155 modulates the interleukin-1 signaling pathway in activated human monocyte-derived dendritic cells. Proc Natl Acad Sci U S A 2009;106:2735-40. [Crossref] [PubMed]
- Wang P, Hou J, Lin L, et al. Inducible microRNA-155 feedback promotes type I IFN signaling in antiviral innate immunity by targeting suppressor of cytokine signaling 1. J Immunol 2010;185:6226-33. [Crossref] [PubMed]
- Lu D, Yamawaki T, Zhou H, et al. Limited differential expression of miRNAs and other small RNAs in LPS-stimulated human monocytes. PLoS One 2019;14:e0214296. [Crossref] [PubMed]
- Goodridge HS, Reyes CN, Becker CA, et al. Activation of the innate immune receptor Dectin-1 upon formation of a 'phagocytic synapse'. Nature 2011;472:471-5. [Crossref] [PubMed]
- Taylor PR, Tsoni SV, Willment JA, et al. Dectin-1 is required for beta-glucan recognition and control of fungal infection. Nat Immunol 2007;8:31-8. [Crossref] [PubMed]
- Agustinho DP, de Oliveira MA, Tavares AH, et al. Dectin-1 is required for miR155 upregulation in murine macrophages in response to Candida albicans. Virulence 2017;8:41-52. [PubMed]
- Netea MG, Brown GD, Kullberg BJ, et al. An integrated model of the recognition of Candida albicans by the innate immune system. Nat Rev Microbiol 2008;6:67-78. [Crossref] [PubMed]
- Ferwerda G, Netea MG, Joosten LA, et al. The role of Toll-like receptors and C-type lectins for vaccination against Candida albicans. Vaccine 2010;28:614-22. [Crossref] [PubMed]
- O'Connell RM, Taganov KD, Boldin MP, et al. MicroRNA-155 is induced during the macrophage inflammatory response. Proc Natl Acad Sci U S A 2007;104:1604-9. [Crossref] [PubMed]
- Yin Q, Wang X, McBride J, et al. B-cell receptor activation induces BIC/miR-155 expression through a conserved AP-1 element. J Biol Chem 2008;283:2654-62. [Crossref] [PubMed]
- Lali FV, Hunt AE, Turner SJ, et al. The pyridinyl imidazole inhibitor SB203580 blocks phosphoinositide-dependent protein kinase activity, protein kinase B phosphorylation, and retinoblastoma hyperphosphorylation in interleukin-2-stimulated T cells independently of p38 mitogen-activated protein kinase. J Biol Chem 2000;275:7395-402. [Crossref] [PubMed]
- Taganov KD, Boldin MP, Chang KJ, et al. NF-kappaB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc Natl Acad Sci U S A 2006;103:12481-6. [Crossref] [PubMed]
- Migita K, Iwanaga N, Izumi Y, et al. TNF-alpha-induced miR-155 regulates IL-6 signaling in rheumatoid synovial fibroblasts. BMC Res Notes 2017;10:403. [Crossref] [PubMed]
- Zhu M, Wang M, Yang F, et al. miR-155-5p inhibition promotes the transition of bone marrow mesenchymal stem cells to gastric cancer tissue derived MSC-like cells via NF-kappaB p65 activation. Oncotarget 2016;7:16567-80. [PubMed]
- Wu XY, Fan WD, Fang R, et al. Regulation of microRNA-155 in endothelial inflammation by targeting nuclear factor (NF)-kappaB P65. J Cell Biochem 2014;115:1928-36. [PubMed]
- Wang M, Yang F, Qiu R, et al. The role of mmu-miR-155-5p-NF-kappaB signaling in the education of bone marrow-derived mesenchymal stem cells by gastric cancer cells. Cancer Med 2018;7:856-68. [Crossref] [PubMed]
- Jia XM, Tang B, Zhu LL, et al. CARD9 mediates Dectin-1-induced ERK activation by linking Ras-GRF1 to H-Ras for antifungal immunity. J Exp Med 2014;211:2307-21. [Crossref] [PubMed]


