Real-time liquid biopsies become a reality in cancer treatment
Certain fragments of DNA shed by tumours into the bloodstream can potentially be used to non-invasively screen for early-stage cancers, monitor responses to treatment and help explain why some cancers are resistant to therapies. For most tumours, a tissue biopsy is quite challenging in that it is costly, painful, or potentially risky for the patient. All these are good reasons to learn about cancer through blood and to get excited about the possibility of carrying out liquid biopsies. The development of non-invasive methods to detect and monitor tumours continues to be a major challenge in oncology. Cell-free circulating tumour DNA (ctDNA) and circulating tumour cells (CTCs) are plasma sources of tumour DNA that have been investigated for non-invasive detection and monitoring of patient tumours but have not been analysed or directly compared across multiple tumour types. Although the current Food and Drug Administration (FDA)-approved liquid biopsy measures intact CTCs to give a prognosis of overall survival, the potential predictive value of ctDNA is much more exciting. ctDNA liquid biopsy allows us to understand specifically what kind of molecular changes are happening in the tumour in real time, which is a very big step beyond where CTCs are today in clinical terms.
An interesting study in this field, providing a wealth of information on the potential utility of ctDNA measurements to assess patients with various cancers, was published in the February 19 issue of Science Translational Medicine (1). The aim of this study was to measure levels of ctDNA with tumour-specific DNA mutations or structural changes in large cohorts of matched human plasma and tumour tissue samples. Bettegowda et al., used a sophisticated method of digital polymerase chain reaction-based sequencing techniques and rearrangement analyses to measure levels of ctDNA with tumour-specific DNA mutations or structural changes in large cohorts of matched human plasma and tumour tissue samples (1). Among study participants with metastatic cancers, ctDNA was detected in 82% of patients with solid tumours outside the brain, including more than 75% of patients with advanced ovarian, colorectal, bladder, gastroesophageal, pancreatic, breast, hepatocellular, and head and neck cancers, as well as melanomas. Less than 50% of patients with medulloblastomas or metastatic cancers of the kidney, prostate or thyroid, and less than 10% of patients with gliomas, had detectable ctDNA (1). Among patients with localized cancer of all types evaluated, only 55% had detectable ctDNA, but the proportion of patients with detectable ctDNA increased with tumour stage (1) (Table 1). Of note, CTCs were often absent in patients with detectable ctDNA; when CTCs and ctDNA were present in the same patient, the average number of mutated DNA fragments was greater in ctDNA isolations, highlighting the distinct nature of these biomarkers (1).

Full table
Early detection is the holy grail of cancer research and any effort to enhance detection of cancer at an early stage is more than welcome. Recently, Ilie et al., reasoned that the invasive character of lung cancer could be used as its “Achilles’ heel” and demonstrated the proof of concept of CTC detection as an early indicator of invasive lung cancer in “at risk” patients. They examined the presence of CTCs by an International Symposium on Endovascular Therapy (ISET) filtration-enrichment technique in complement to CT-scan in chronic obstructive pulmonary disease (COPD) patients without clinically detectable lung cancer (3). CTCs were able to be detected in patients with COPD and monitoring “sentinel” CTC-positive COPD patients allowed early diagnosis of lung cancer (3) (Table 1).
Bettegowda and colleagues were able to detect circulating KRAS-mutant DNA fragments in the plasma of patients with KRAS-mutant tumours with high specificity and sensitivity, and high ctDNA levels were associated with decreased 2-year survival. A separate panel of 206 patients with metastatic colorectal cancer was examined to determine the sensitivity and specificity of ctDNA liquid biopsies. The sensitivity of ctDNA for detection of clinically relevant KRAS gene mutations was 87.2% and its specificity was 99.2% (Table 1). In patients with metastatic colorectal cancer who developed resistance to EGFR antibodies, analysis of ctDNA identified the emergence of polyclonal KRAS, NRAS, BRAF, or EGFR mutations in 96% of panitumumab- or cetuximab-refractory patients. Subsequently, Misale et al., were able to illustrate a way to use this information to overcome treatment resistance (2). Regardless of the genetic alterations, resistant cells consistently displayed MEK and ERK activation, which persisted after EGFR blockade, while suppression of EGFR, together with silencing of MEK, was required to hamper the proliferation of resistant cells (2). Therefore MEK inhibitors, in combination with cetuximab or panitumumab, should be tested in Colorectal Cancer (CRC) patients who become refractory to anti-EGFR therapies (Table 1).
Besides DNA, tumour cells can also release RNA into the blood by a variety of microvesicle-dependent or -independent mechanisms (5,6). Several models have shown that platelets are crucial to tumour cell growth, spread, invasion, intravasation, migration, extravasation, and establishment of distant metastasis (7,8). Cancer cells have a remarkable arsenal of mechanisms to induce platelet aggregation (8). Tumour cells release tumour-derived biomolecules into the blood, where they are absorbed by platelets (4,9-12). Using confocal microscopy and reverse transcription-PCR (RT-PCR), it has been shown that platelets isolated from healthy individuals take up RNA-containing membrane vesicles from cancer cells (4). Moreover, platelets isolated from patients with glioma and prostate cancer contained the cancer-associated RNA biomarkers EGFRvIII and prostate cancer antigen 3 (PCA3), respectively (4) (Table 1). Since platelet RNA can be readily isolated and analyzed (13,14) platelets seem to be an additional source for the non-invasive monitoring of biomarkers. Furthermore, microvesicles, like exosomes, are present in plasma and may provide a protective environment for tumour-derived RNA though specialized techniques are required for adequate RNA isolation (5) (Figure 1).
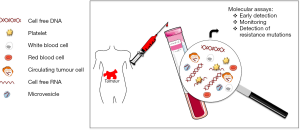
To date, liquid biopsies have generated a lot of excitement since they can provide a non-invasive, ongoing picture of a patient’s cancer, offering valuable insight into how best to fight it. In addition to offering clues about stage and spread, liquid biopsies can be used to monitor the effects of cancer treatment, give an early warning about possible recurrence and offer clues to the reasons for treatment resistance. ctDNA investigations in cancer patients are increasingly being performed, supporting the different potential applications of this approach. Serial analysis of ctDNA during treatment can provide a dynamic picture of molecular disease changes, suggesting that this noninvasive approach could also be used to monitor the development of secondary resistance and identify heterogeneous subclonal populations of tumour cells developing during the course of treatment. In the future, instead of extensive imaging and invasive tissue biopsies, liquid biopsies could be used to guide cancer treatment decisions and perhaps even screen for tumours that are not yet visible on imaging.
Acknowledgements
Disclosure: The authors declare no conflict of interest.
References
- Bettegowda C, Sausen M, Leary RJ, et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med 2014;6:224ra-24. [PubMed]
- Misale S, Arena S, Lamba S, et al. Blockade of EGFR and MEK intercepts heterogeneous mechanisms of acquired resistance to anti-EGFR therapies in colorectal cancer. Sci Transl Med 2014;6:224ra-26. [PubMed]
- Ilie M, Hofman V, Long-Mira E, et al. "Sentinel" circulating tumor cells allow early diagnosis of lung cancer in patients with chronic obstructive pulmonary disease. PLoS One 2014;9:e111597. [PubMed]
- Nilsson RJ, Balaj L, Hulleman E, et al. Blood platelets contain tumor-derived RNA biomarkers. Blood 2011;118:3680-3. [PubMed]
- Skog J, Würdinger T, van Rijn S, et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat Cell Biol 2008;10:1470-6. [PubMed]
- Nilsson J, Skog J, Nordstrand A, et al. Prostate cancer-derived urine exosomes: a novel approach to biomarkers for prostate cancer. Br J Cancer 2009;100:1603-7. [PubMed]
- Gay LJ, Felding-Habermann B. Contribution of platelets to tumour metastasis. Nat Rev Cancer 2011;11:123-34. [PubMed]
- Goubran HA, Stakiw J, Radosevic M, et al. Platelets effects on tumor growth. Semin Oncol 2014;41:359-69. [PubMed]
- Klement GL, Yip TT, Cassiola F, et al. Platelets actively sequester angiogenesis regulators. Blood 2009;113:2835-42. [PubMed]
- Labelle M, Begum S, Hynes RO. Direct signaling between platelets and cancer cells induces an epithelial-mesenchymal-like transition and promotes metastasis. Cancer Cell 2011;20:576-90. [PubMed]
- McAllister SS, Weinberg RA. The tumour-induced systemic environment as a critical regulator of cancer progression and metastasis. Nat Cell Biol 2014;16:717-27. [PubMed]
- Schumacher D, Strilic B, Sivaraj KK, et al. Platelet-derived nucleotides promote tumor-cell transendothelial migration and metastasis via P2Y2 receptor. Cancer Cell 2013;24:130-7. [PubMed]
- Goodall AH, Burns P, Salles I, et al. Transcription profiling in human platelets reveals LRRFIP1 as a novel protein regulating platelet function. Blood 2010;116:4646-56. [PubMed]
- Nagalla S, Shaw C, Kong X, et al. Platelet microRNA-mRNA coexpression profiles correlate with platelet reactivity. Blood 2011;117:5189-97. [PubMed]


