A potentially effective drug for patients with recurrent glioma: sermorelin
Introduction
Glioma is the most common primary tumor of the central nervous system (1). Although with the improvement of medical technology, the tumor could be nearly totally resected, and post-operative supplemented with a series of comprehensive treatment such as radiotherapy and chemotherapy (2). But the prognosis of glioma patients is still poor due to the recurrence of tumors. The only first-line drug for clinical treatment of primary glioma is temozolomide (TMZ) (3). As few drugs can be used, the tumor inevitably developed resistance to TMZ and relapsed (4). Recurrence of therapeutically resistant tumors will occur quickly and cause death. Therefore, it is an urgent need to find effective drugs for recurrent gliomas.
There have been some studies to screen effective drugs for recurrent gliomas. Our previous study found that a small molecule drug called Bozitinib which was targeting the PTPRZ1-MET fusion gene in patients with secondary glioblastoma was processing in Phase II clinical trial. It is expected to improve the prognosis of patients with the fusion gene (5). The current research was mostly limited to a few patients with certain clinical characteristics, but most recurrent patients still have no effective drugs (6). Therefore, how to use the existing marketed drugs to treat patients with recurrent glioma became an achievable and convenient method. In this study, 4,865 drugs and the CGGA databases containing sequencing information of primary and recurrent gliomas patients were used to high-throughput drug screening for recurrent gliomas. Finally, we found that sermorelin was the most effective drug for recurrent gliomas.
Sermorelin is a synthetic growth hormone releasing hormone (GHRH) composed of 29 amino acids with a relative molecular weight of 3,357.88. It is the amino-terminal fragment of endogenous GHRH and has a significant effect on regulating growth (7). GHRH is a hypothalamic peptide neurohormone that regulates the release of growth hormone (GH) by the pituitary. It is an autocrine/paracrine growth factor that affects cells by acting on the growth hormone releasing hormone receptor (GHRHR) (8). As a common mediator of the nervous system, immune system and endocrine system, it can not only stimulate pancreatic β cell proliferation and cardiomyocyte survival, but also inhibit tumor cell growth (9).
Jaszberenyi et al. reported that GHRH agonist combined with doxorubicin treatment can significantly inhibit the growth, invasion, and metastasis of glioblastoma cell line U-87 in vitro and in vivo (10). Schally et al. also reported that GHRH agonists inhibit the growth of tumors by down-regulating the expression of GHRHR in vivo tests (11). Based on the analysis of sequencing results of glioma samples from CGGA databases, we found that there is a higher expression of GHRHR in the glioma samples of recurrent patients, suggesting that sermorelin may suppress the tumor growth by down-regulating the expression of GHRHR and glial growth factor (GGF).
Taking the advantages of the CGGA database, which has the sequencing and follow-up data of patients with recurrent gliomas, we screened out the most effective drug for recurrent gliomas patients. Through further analysis, we found that sermorelin is more suitable for patients with much malignant molecular phenotypes, including WHO grade IV, IDH wildtype, 1p/19q non-codeletion, and mesenchymal subtype gliomas. Gene ontology (GO) and Kyoto encyclopedia of genes and genomes (KEGG) analyses found that the Drug Resistance Score (DRS) of sermorelin was negatively correlated with cell proliferation and immune function. Our previous studies showed that in patients with glioblastoma, higher immune cell infiltration corresponds to a worse prognosis (12). After the analysis of immune checkpoints and immune cell components, we found that patients with low DRS had higher immune checkpoint expression and macrophage infiltration, suggesting that these patients were more sensitive to sermorelin. Finally, we analyzed the prognosis of DRS and patients receiving radiotherapy and chemotherapy. We found that patients with low DRS had a worse prognosis. Moreover, patients who had received radiotherapy and chemotherapy have lower DRS. It means that these treated patients were more sensitive to sermorelin and may benefit more by using sermorelin. Our study was the first time to use large sample sequencing analysis to find available drugs for patients with recurrent gliomas after applying all the existing treatments. We present the following article in accordance with the MDAR checklist (available at http://dx.doi.org/10.21037/atm-20-6561).
Methods
Data collection
The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). This study was approved by the Capital Medical University Institutional Review Board (Number: KY2014-002-02) and wrote informed consent was obtained from each patient. We collected transcriptome sequencing data generated by Illumina HiSeq platform from CGGA database. Among CGGA database, 325 samples transcriptome sequencing data were completed in 2016, while other 693 samples data were completed in 2019. Thus, these two sets of data were completed respectively at different time and considered as independent. In this study, we used 325 samples as discovery cohort and 693 samples as validation cohort. Clinical and molecular information was obtained from the CGGA website (https://www.cgga.org.cn). All clinical and molecular information of the samples is presented in Table 1. The information of 4,865 drugs were downloaded from DrugBank online (https://go.drugbank.com/) (13). Our previous study had described the method to detect IDH mutation status and 1p/19q co-deletion status (14,15). 15 and 23 pair-validated patient samples were retrieved from the GSE62153 and GSE4271 databases of the Gene Expression Omnibus database, respectively.
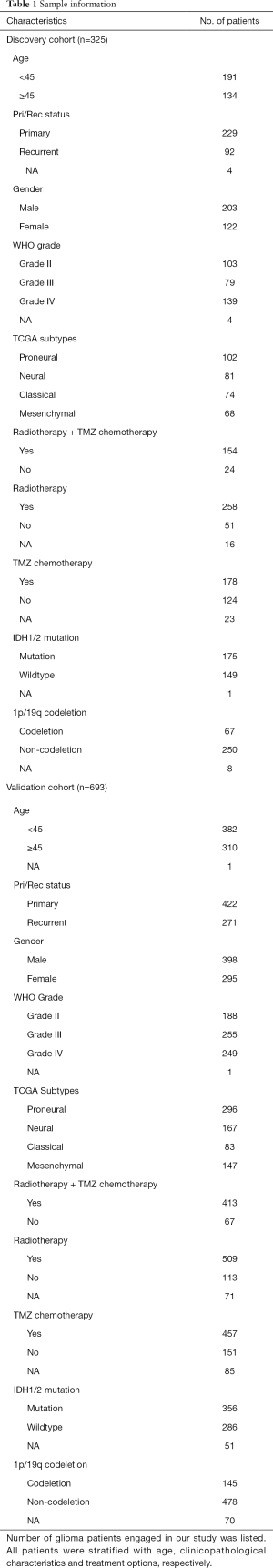
Full table
Chemicals and reagents
Sermorelin (purchased from ACMEC biochemical, CAS: 86168-78-7). DMEM basic medium (Gibco). Fetal bovine serum (Gibco). Dimethyl sulfoxide (Sigma). Penicillin-Streptomycin (Gibco). Cell Counting Kit-8 (Dojindo).
Cell culture
The human glioblastoma cell lines U87 and LN229 were purchased from National Infrastructure of Cell Line Resource (http://www.cellresource.cn/). Cells were cultured in DMEM basic medium supplemented with 10% fetal bovine serum and 1% Penicillin-Streptomycin, incubated at 37 °C with 5% CO2.
Cell counting kit-8 (CCK-8) assay
Cells incubated in 96-well plates were treated as indicated and cell proliferation was assessed by CCK-8 assay at 24 and 48 h post treatment following the manufacturer’s instruction. Optical density (OD) was recorded at 450 nm.
Gene Set Variation Analysis (GSVA)
GSVA was performed with the ‘GSVA’ package (from the R Project 3.5.1) of R software. Default parameters were used in GSVA analysis and the gene list of each drug was downloaded from the Bader Lab geneset (https://www.baderlab.org).
Gene ontology (GO) and Kyoto encyclopedia of genes and genomes (KEGG) analyses
The biological functions and signaling pathways related to sermorelin were explored by GO and KEGG analyses using the DAVID bioinformatics resource (version 6.7) (16).
Biological function analysis
Biological function scores were calculated by GSVA analysis and each biological function geneset was downloaded from AmiGO 2 Web portals (http://amigo.geneontology.org/amigo/landing).
CIBERSORT
RNA-seq data from CGGA databases were evaluated using the CIBERSORT software (https://cibersort.stanford.edu). The signature gene profile of 22 immune cell types was used in CIBERSORT to estimate the proportion of tumor-infiltrating immune cell types (17).
Prognostic analysis
Patient survival distribution and significance were evaluated by the Kaplan-Meier survival curve and log-rank test. Kaplan-Meier analysis was performed using R software (version 3.5.1, http://www.r-project.org). The prognostic value of the DRS was estimated by univariate and multivariate Cox proportional hazard model analysis using SPSS statistical software (version 25.0; IBM, Armonk, NY, USA). Patients with missing information were excluded from the analysis.
Statistical analysis
The Student t-test, one-way ANOVA, or Chi-squared test was used to assess differences in variables between groups. Other statistical computations and figures drawing were performed with several packages (ggplot2, survival, and corrgram) in the statistical software environment R, version 3.3.2 (https://www.r-project.org), and Graphpad Prism 7 (https://www.graphpad.com). For all statistical methods, P<0.05 was considered as significant difference.
Results
Sermorelin is the most effective drug to recurrent glioma patients
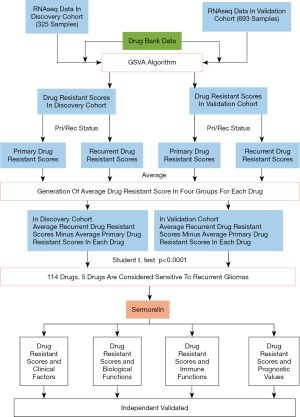
To find the most effective drugs for patients with recurrent glioma among the drugs which have been approved by the FDA (the U.S. Food and Drug Administration), CGGA databases including the RNA sequencing data of each patient was been used. The specific work flow was shown in Figure 1. At first, we used GSVA analysis to calculate the DRS based on the unique drug resistant genes of each drug. 4,865 drugs were put into analysis and each drug was generated a DRS for each patient. Next, we divided the DRS into two groups according to whether the patients were primary or recurrent glioma of each drug. Lastly, we calculated the average value of two groups respectively and each drug was generated a recurrent score and a primary score. We defined the difference value calculated by recurrent score minus primary score as the symbol of drug sensitivity. A positive value means drug resistance and a negative value means drug sensitivity in two cohorts (Figure 2A,B). After comparing them with the Student t-test, we found 2.34% (114/4,865) of these drugs had statistical difference in two cohorts. According to these values, we found only five drugs, which had the 5 largest difference value (P<0.0001), and sermorelin had the lowest p-value and maximum difference value among these drugs in both discovery and validation cohorts (Figure 2C). GHRHR was the most decisive gene of the sensitivity of glioma to sermorelin. Fifteen and 23 patients from GSE62153 and GSE4271 databases were recurrent glioma patients, respectively. Meanwhile, they had both primary gliomas and recurrent gliomas RNA-seq data. We compared two status and found that GHRHR expression increased in the recurrent gliomas (Figure 2D,E). To further comfired whether sermorelin could inhibit glioma cell growth, CCK8 assay was performed to evaluate the results in U87 and LN229 cell lines. We found that sermorelin inhibited glioma cell growth in a dose- and time-dependent manner (Figure 2F,G). Above all, we considered sermorelin was the most effective drug for recurrent glioma patients and the following research was focus on it.

The landscape of DRS and clinical characteristics in glioma
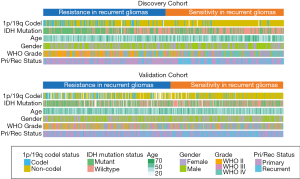
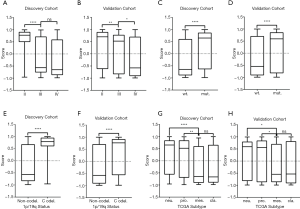
According to the DRS calculated by the GSVA analysis of sermorelin, we divided patients into the resistant group and the sensitive group from both discovery and validation cohorts. The median of the scores was setting as a cutoff. Based on the GSVA analysis we defined the specimens with scores less than the median as sensitive to sermorelin and the other parts were resistance ones. Heatmap showed a comparison between two groups in the fields including WHO grade, age, gender, IDH status, 1p/19q status, and primary/recurrent status. Scrutinizing the graph, except for gender and age, each of the other characteristics had unsymmetrical distribution. Higher-grade, IDH-wildtype status, 1p/19q non-codeletion status, recurrent glioma patients were more frequently distributed in the sensitive group (Figure 3). The correlation between DRS of sermorelin and various clinical features were inquired in discovery and validation cohorts. We found DRS in WHO grade IV was lower than WHO grade II gliomas in discovery cohort. A similar trend could also be found in validation cohort (Figure 4A,B). The IDH mutation status and 1p/19q co-deletion act as important indicators for prognosis of gliomas. We found the DRS was much lower in IDH-wildtype and 1p/19q non-codeletion gliomas in both cohorts (Figure 4C,D,E,F). Besides, we investigated DRS in different molecular subtypes defined by TCGA network (18). Among transcriptome subtypes, the scores were relatively lower in the Mesenchymal and Classical subtypes compared with other two subtypes in both cohorts (Figure 4G,H). Acknowledgedly, higher-grade, IDH-wildtype, and 1p/19q non-codeletion status mean a poor prognosis and an insensitive treatment response. But interestingly, our findings revealed that these refractory patients were more sensitive to sermorelin.
Sermorelin is closely related to cell proliferation functions
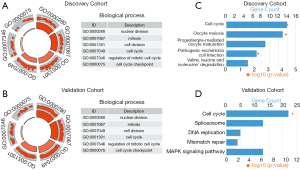
To clarify the biologic role of sermorelin in glioma, we performed GO and KEGG analyses. Firstly, we created a gene list that strongly correlated with sermorelin by Pearson correlation analysis (Pearson|R|>0.45 and P<0.0001). Then we chose the most related 500 genes in both discovery and validation cohorts gene lists. Lastly, we explored the biological function of these genes respectively by GO analysis in DAVID Bioinformatics Resources 6.7. When the gene functions were sorted by P value in increasing order, genes most relevant to sermorelin were mostly involved in cell proliferation function, like nuclear division and cell cycle, in both discovery and validation cohorts (Figure 5A,B). Additionally, we performed KEGG pathway analysis to further explore the signaling pathways associated with the above-mentioned genes. Expectedly, the KEGG analysis identified these genes to be associated with cell cycle pathway in two cohorts (Figure 5C,D). We concluded sermorelin may inhibit glioma cell proliferation by blocking cell cycle progression or preventing nuclear division.
Sermorelin is related to immune system
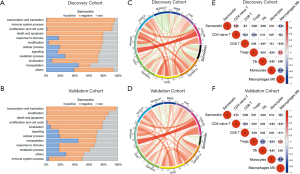
To further investigate the biological function of sermorelin, we put 7,350 biological functions, which were confirmed by GO analysis in AmiGO 2 website, into GSVA analysis and biological scores were generated (19). We compared them with the DRS of sermorelin and Pearson correlation analysis was been used to find the most related functions with sermorelin. In both cohorts, we found sermorelin was related to “transcription and translation, immune system process, proliferation and cell cycle” (Figure 6A,B). It further verified the conclusion that sermorelin blocked cell cycle progression to inhibit cell proliferating. Moreover, we found sermorelin had a strong association with the immune system. To further explore the role of sermorelin in the immune response in glioma, we adopted the Pearson correlation analysis to investigate whether immune checkpoints were related to sermorelin. The result indicated that most immune checkpoints were negatively related to sermorelin in both cohorts (Figure 6C,D). Then we investigated which types of immune cells contributed to this process. We estimated the abundance of infiltrated immune cells by CIBERSORT and found that CD4 naïve T cells and monocytes had a positive correlation with sermorelin while macrophages M0 was opposite in discovery and validation cohorts (Figure 6E,F). This finding demonstrated that besides modulating cell proliferation, sermorelin played a vital role in glioma through regulating the immune system.
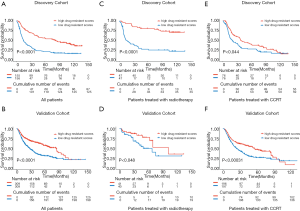
Patients with poorer prognoses are more sensitive to sermorelin
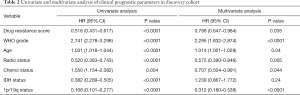
In consideration of the robust negative relationship between sermorelin and immune status, we analyzed the prognostic value of sermorelin in both cohorts. We analyzed the prognoses of a total of 929 glioma patients by the Kaplan-Meier method. In both cohorts, the patients with lower DRS of sermorelin had worse prognoses (Figure 7A,B). It verified patients with poor prognoses benefited more from using sermorelin. Interestingly, our study found that, for patients who had been treated with radiotherapy or concurrent chemoradiotherapy (CCRT), the lower DRS of sermorelin conferred to poorer prognoses (Figure 7C,D,E,F). It means that these treated patients were more sensitive to sermorelin. Furthermore, uni- and multivariate Cox regression analysis were performed, and the independence of the clinical prognostic significance of DRS of sermorelin in glioma was verified. In both cohorts, it was shown that DRS, WHO Grade, age at diagnosis, radiotherapy status, chemotherapy status, and 1p/19q status were significantly associated with overall survival. In multivariate analysis, the DRS was also a significant factor after adjusting for the clinical factors mentioned above (Tables 2,3). These findings indicated that patients with poor prognoses were sensitive to sermorelin.
Full table
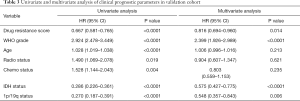
Full table
Discussion
Glioma patients generally had poor prognoses, especially those with tumors recurrent after treatment with temozolomide (20). Due to the resistance to temozolomide, effective chemotherapy cannot be achieved after surgery, resulting in dismal outcomes of patients with recurrent glioma (21). Temozolomide was used as an “orphan drug” in the chemotherapy of gliomas so far. Moreover, rigorous and long-term clinical trials were needed in the development of new drugs. Therefore, exploring the drugs had the potential to treat recurrent glioma patients with diverse drug-screening approaches, became a simple and quick approach. The current drug screening approach was only applicable to individual patients. The typical approaches include implanting glioma tissues removed by neurosurgery into mice subcutaneously or orthotopically to construct a PDX (patient-derived xenograft) model or employing tumor cell lines capable of stable passage as models for drug screening (22,23). These approaches were time-consuming and costing long culture periods. Because of the strong heterogeneity of gliomas, the applications of the drugs screened by the above approach were limited to a few patients. Most patients could not benefit from it. Our research provides a high-throughput screening method for drug sensitivity based on transcriptomics. It is also the first research to find drugs that are sensitive for the majority population of recurrent glioma patients by combining large-scale samples RNA sequencing and clinical characteristics of patients. Before the development of new drugs, the survival time of recurrent glioma patients could be extended as much as possible by using sermorelin.
We acquired the drug sermorelin via the above analysis. As a GHRH analog, it stimulates GHRHR to play a regulatory role. Sermorelin could penetrate the blood-brain barrier easily accompanied by fewer side effects. It is ideal for treating patients with recurrent glioma (24,25). Interestingly, our analysis also found that patients with stronger malignant phenotypes were more sensitive to sermorelin. Patients with gliomas of IDH wildtype, 1p/19q non-codeletion, and mesenchymal subtype had worse prognoses and shorter average survival time. The DRS of sermorelin was lower in these patients meaning that these patients were more sensitive to sermorelin. Patients in this category will benefit more from treatment with sermorelin. Meanwhile, our analysis also revealed that sermorelin was related to immune function. The stronger enrichments of immune checkpoints features and M0 macrophages, and the lower DRS of the sermorelin means the greater sensitivity to sermorelin. We speculated that sermorelin could withstand the depletion of immune components caused by tumor invasion, increase the components of immune cells, thereby enhancing the function of the immune system and benefiting patients (26).
As an analogue of GHRH, sermorelin could inhibit the proliferation of tumor cells similarly to most other chemotherapeutic drugs (27-29). Our analysis revealed that sermorelin could inhibit transcription and translation of tumor cells by regulating the cell cycle and nuclear division. Malignant tumors grow fast due to their rapid propagation capability and thus are more sensitive to sermorelin. The previous study demonstrated that the combination of GHRH agonist JI-34 and doxorubicin can inhibit the growth of glioma cell line U-87 by reducing the expression of apoptosis genes in vivo and in vitro (10). Schally et al. also found that GHRH agonist MR409 inhibited the growth of various human-derived tumor cells by down-regulating GHRHR (11). Based on this, we believed that GHRH agonists could benefit recurrent glioma patients in a way of using alone or in combination with TMZ.
Our research provided a new idea and approach for cancer drug screening. Although individualized drug sensitivity screening can provide patients with suitable drugs accurately, this will be a long process (30,31). For recurrent glioma patients, using the existing large-scale samples sequencing data to find effective drugs for patients could save time and effort. What’s more, our research also provided new insights for the chemotherapy of other tumors. Similar data analysis algorithms could be extended to pancreatic cancer, small cell lung cancer, lymphoma, and other cancers. Combining their clinical characteristics and pathological characteristics to explore effective drugs brings us new hopes for clinical cancer treatments.
Acknowledgments
We thank Ms. Shuqing Sun and Hua Huang for tissue sample collection and clinical data retrieval.
Funding: This work was supported by grants from the National Natural Science Foundation of China (No. 81972816, 81672479, 81903078), Construction of the Genomics Platform for Chinese People’s Brain Diseases (No. PXM2019_026280_000002), Beijing Nova Program (Z201100006820118).
Footnote
Reporting Checklist: The authors have completed the MDAR checklist. Available at http://dx.doi.org/10.21037/atm-20-6561
Data Sharing Statement: Available at http://dx.doi.org/10.21037/atm-20-6561
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/atm-20-6561). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). This study was approved by the Capital Medical University Institutional Review Board (Number: KY2014-002-02) and wrote informed consent was obtained from each patient.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Van Meir EG, Hadjipanayis CG, Norden AD, et al. Exciting new advances in neuro-oncology: the avenue to a cure for malignant glioma. CA Cancer J Clin 2010;60:166-93. [Crossref] [PubMed]
- Jiang T, Mao Y, Ma W, et al. CGCG clinical practice guidelines for the management of adult diffuse gliomas. Cancer Lett 2016;375:263-73. [Crossref] [PubMed]
- Herrlinger U, Tzaridis T, Mack F, et al. Lomustine-temozolomide combination therapy versus standard temozolomide therapy in patients with newly diagnosed glioblastoma with methylated MGMT promoter (CeTeG/NOA-09): a randomised, open-label, phase 3 trial. Lancet 2019;393:678-88. [Crossref] [PubMed]
- Chai RC, Zhang KN, Liu YQ, et al. Combinations of four or more CpGs methylation present equivalent predictive value for MGMT expression and temozolomide therapeutic prognosis in gliomas. CNS Neurosci Ther 2019;25:314-22. [Crossref] [PubMed]
- Hu H, Mu Q, Bao Z, et al. Mutational Landscape of Secondary Glioblastoma Guides MET-Targeted Trial in Brain Tumor. Cell 2018;175:1665-78 e18.
- van den Bent MJ, Klein M, Smits M, et al. Bevacizumab and temozolomide in patients with first recurrence of WHO grade II and III glioma, without 1p/19q co-deletion (TAVAREC): a randomised controlled phase 2 EORTC trial. Lancet Oncol 2018;19:1170-9. [Crossref] [PubMed]
- Prakash A, Goa KL. Sermorelin - A review of its use in the diagnosis and treatment of children with idiopathic growth hormone deficiency. Biodrugs 1999;12:139-57. [Crossref] [PubMed]
- Walker RF. Sermorelin: A better approach to management of adult-onset growth hormone insufficiency? FORE-WORD. Clin Interv Aging 2006;1:307-8. [Crossref] [PubMed]
- Sinha DK, Balasubramanian A, Tatem AJ, et al. Beyond the androgen receptor: the role of growth hormone secretagogues in the modern management of body composition in hypogonadal males. Transl Androl Urol 2020;9:S149-59. [Crossref] [PubMed]
- Jaszberenyi M, Rick FG, Popovics P, et al. Potentiation of cytotoxic chemotherapy by growth hormone-releasing hormone agonists. Proc Natl Acad Sci U S A 2014;111:781-6. [Crossref] [PubMed]
- Kiaris H, Schally AV. Decrease in telomerase activity in U-87MG human glioblastomas after treatment with an antagonist of growth hormone-releasing hormone. Proc Natl Acad Sci U S A 1999;96:226-31. [Crossref] [PubMed]
- Wang ZL, Wang Z, Li GZ, et al. Immune Cytolytic Activity Is Associated With Genetic and Clinical Properties of Glioma. Front Immunol 2019;10:1756. [Crossref] [PubMed]
- Wishart DS, Feunang YD, Guo AC, et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res 2018;46:D1074-82. [Crossref] [PubMed]
- Chai RC, Zhang KN, Chang YZ, et al. Systematically characterize the clinical and biological significances of 1p19q genes in 1p/19q non-codeletion glioma. Carcinogenesis 2019;40:1229-39. [Crossref] [PubMed]
- Wang HY, Tang K, Liang TY, et al. The comparison of clinical and biological characteristics between IDH1 and IDH2 mutations in gliomas. J Exp Clin Cancer Res 2016;35:86. [Crossref] [PubMed]
- Huang DW, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bio-informatics resources. Nat Protoc 2009;4:44-57. [Crossref] [PubMed]
- Newman AM, Steen CB, Liu CL, et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat Biotechnol 2019;37:773. [Crossref] [PubMed]
- Verhaak RGW, Hoadley KA, Purdom E, et al. Integrated Genomic Analysis Identifies Clinically Relevant Subtypes of Glioblastoma Characterized by Abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 2010;17:98-110. [Crossref] [PubMed]
- Li G, Wang Z, Zhang C, et al. Molecular and clinical characterization of TIM-3 in glioma through 1,024 samples. Oncoimmunology 2017;6:e1328339. [Crossref] [PubMed]
- Gaviani P, Silvani A, Lamperti E, et al. Rechallenge with temozolomide in recurrent glioma. Neurol Sci 2011;32:S247-9. [Crossref] [PubMed]
- Wei W, Chen X, Ma XM, et al. The efficacy and safety of various dose-dense regimens of temozolomide for re-current high-grade glioma: a systematic review with meta-analysis. J Neurooncol 2015;125:339-49. [Crossref] [PubMed]
- Lee JK, Liu ZQ, Sa JK, et al. Pharmacogenomic landscape of patient-derived tumor cells informs precision oncology therapy. Nat Genet 2018;50:1399-411. [Crossref] [PubMed]
- Jacob F, Salinas RD, Zhang DY, et al. A Patient-Derived Glioblastoma Organoid Model and Biobank Recapitulates Inter- and Intra-tumoral Heterogeneity. Cell 2020;180:188-204.e22. [Crossref] [PubMed]
- Jaeger LB, Banks WA, Varga JL, et al. Antagonists of growth hormone-releasing hormone cross the blood-brain barrier: a potential applicability to treatment of brain tumors. Proc Natl Acad Sci U S A 2005;102:12495-500. [Crossref] [PubMed]
- Schally AV, Wang H, He J, et al. Agonists of growth hormone-releasing hormone (GHRH) inhibit human experimental cancers in vivo by down-regulating receptors for GHRH. Proc Natl Acad Sci U S A 2018;115:12028-33. [Crossref] [PubMed]
- Zhang C, Cai R, Lazerson A, et al. Growth Hormone-Releasing Hormone Receptor Antagonist Modulates Lung Inflammation and Fibrosis due to Bleomycin. Lung 2019;197:541-9. [Crossref] [PubMed]
- Wang H, Zhang X, Vidaurre I, et al. Inhibition of experimental small-cell and non-small-cell lung cancers by novel antagonists of growth hormone-releasing hormone. Int J Cancer 2018;142:2394-404. [Crossref] [PubMed]
- Munoz-Moreno L, Schally AV, Prieto JC, et al. Growth hormone-releasing hormone receptor antagonists modify molecular machinery in the progression of prostate cancer. Prostate 2018;78:915-26. [Crossref] [PubMed]
- Koster F, Engel JB, Schally AV, et al. Triple-negative breast cancers express receptors for growth hor-mone-releasing hormone (GHRH) and respond to GHRH antagonists with growth inhibition. Breast Cancer Res Treat 2009;116:273-9. [Crossref] [PubMed]
- Hou S, Tiriac H, Sridharan BP, et al. Advanced Development of Primary Pancreatic Organoid Tumor Models for High-Throughput Phenotypic Drug Screening. SLAS Discov 2018;23:574-84. [Crossref] [PubMed]
- Pemovska T, Bigenzahn JW, Superti-Furga G. Recent advances in combinatorial drug screening and synergy scoring. Curr Opin Pharmacol 2018;42:102-10. [Crossref] [PubMed]